Production of Two Elite Glutinous Rice Varieties by Editing Wx Gene
2019-02-19FeiYunyanYangJieWangFangquanFanFangjunLiWenqiWangJunXuYangZhuJinyanZhongWeigong
Fei Yunyan, Yang Jie, 2, Wang Fangquan, 2, Fan Fangjun, 2, Li Wenqi, 2, Wang Jun, 2, Xu Yang, 2, Zhu Jinyan, 2, Zhong Weigong, 2
Production of Two Elite Glutinous Rice Varieties by EditingGene
Fei Yunyan1, Yang Jie1, 2, Wang Fangquan1, 2, Fan Fangjun1, 2, Li Wenqi1, 2, Wang Jun1, 2, Xu Yang1, 2, Zhu Jinyan1, 2, Zhong Weigong1, 2
(Institute of Food Crops, Jiangsu Academy of Agricultural Sciences / Nanjing Branch of Chinese National Center for Rice Improvement / Jiangsu High Quality Rice Research & Development Center, Nanjing 210014, China; Jiangsu Co-Innovation Center for Modern Production Technology of Grain Crops, Yangzhou University, Yangzhou 225009, China)
Thegene () in rice, which encodes the granule bound starch synthase enzyme, is responsible for amylose synthesis. Glutinous (sticky) rice has little or no amylose that can be used in various applications, such as brewing. In this study, knockout of thegene with CRISPR/Cas9 technology was conducted in two eliterice lines, Huaidao 5 (HD5) and Suken 118 (SK118), aiming to develop elite sticky rice varieties. We achieved six homozygous T0plants with more than 200 bp deletion in thegene, as well as 36-HD5 and 18-SK118 homozygous transgene-free plants in the T1generation. The seeds of all the mutants were white and opaque, similar to those of sticky rice, and contained only 2.6%–3.2% amylose. Results of scanning electron microscopy showed that the quality of rice did not change. In conclusion, we successfully developed two elite sticky rice varieties.
amylose; CRISPR; Cas9; rice; starch;
Rice is one of the most important crops globally. High yield, good quality and resistant traits are important targets in its breeding programs (Kiswara et al, 2014), and genetic diversity in rice germplasm is crucial for these purposes. Various strategies, such as exploring wild rice germplasm and natural or artificial mutation, have been employed by breeders to broaden the genetic diversity of rice (Shen et al, 2017). However, these methods are time consuming, and in the case of gene mutations, randomly. The clustered regularly interspaced short palindromic repeats/CRISPR-associated 9 (CRISPR/Cas9) system is a novel technology, widely used in various fields because of its cost effective, simple, flexible and highly efficient (Wang et al, 2017a). In rice, numerous studies have been performed with the CRISPR/Cas9 system (Wang et al, 2016). Ma et al (2015) reported that editing efficiency of CRISPR/Cas9 is higher in rice than in. Mikami et al (2015) found that thepromoter performs better than thein mutation frequency. A 1-bp mismatch of an sgRNA is sufficient to avoid off-target in closely related genes in rice (Baysal et al, 2016). There is a greater probability of large gene deletions inrice than inrice (Wang et al, 2017b). With the rapid development of the CRISPR-Cas system, many researchers have begun to use this technology to improve agronomic traits in rice. For example, rice blast resistance lines are produced by CRISPR/Cas9-targeted mutagenesis of(Wang et al, 2016). Hybrid-compatible lines are developed by knocking out thegenes (Xie et al, 2017). Low-Cs+/Cd rice strains are also created by knocking out the corresponding metal transporter genes (Nieves-cordones et al, 2017; Tang et al, 2017).
Amylose is a linear polymer composed of glucose units, linked by α-D-(1-4) glycosidic linkages. In rice, amylose content (AC) is a complex trait, influenced by multiple genes, mainly the() gene, and the environment (Chen et al, 2008). Thegene encodes the granule-bound starch synthase I (GBSSI), which catalyzes the synthesis of amylose. The expression ofis modulated by many factors, which can lead to a change in AC (Wang et al, 2015). Wu et al (2015) reported that dozens ofgenes regulate the splicing efficiency of. A tetratricopeptide domain-containing protein (flo2) regulates the expression of(Wu et al, 2015).,,,and, as transcription factors, modifyexpression (Wamnugu et al, 2017). Many QTLs, such as, have been identified as genes controlling AC (Fasahat et al, 2014). Glutelin, gibberellin receptor, ADP- glucose pyrophosphorylase and pullulanase also play minor roles in affecting AC (Wamnugu et al, 2017). Here, we used CRISPR/Cas9 technology to edit thegene and generate two elite glutinous rice varieties that could meet the demands of consumers and enrich the rice genetic resources.
Materials and methods
Rice materials and growth conditions
Rice () inbred lines Huaidao 5 (HD5) and Suken 118 (SK118) were used for-mediated co-cultivationtransformation experiments, as described previously (Zheng et al, 2016). During growing seasons, all the materials were cultivated in the greenhouse of the Jiangsu Academy of Agricultural Sciences in Nanjing, Jiangsu Province, China.
Plasmid construction
Thegene (LOC_Os06g04200) was the target of Cas9 endonuclease. According to the principles outlined in Ma et al (2015), two sgRNAs (T1 and T2) were designed on the first extron of thegene from Nipponbare. The sgRNAs were assembled into one vector pYLCRISPR/Cas9-MH usingI (New England Biolabs, Country) and T4 ligase (New England Biolabs, Country). T1 and T2 were driven byandpromoters, respectively. The vector was transformed into thestrain DH5α, and the insertions were verified by sequencing. The primers sequences of the sgRNAs are listed in Table 1. A vector map was generated using vector NTI (Lu and Moriyama, 2004).
Detection of mutations
The genomic DNA of transgenic plants was isolated from rice leaves using the CTAB method (Murray and Thompson, 1980). The PCRs were conducted using a Golden Star T6 Super PCR Mix (TsingKe Biotech Co., Ltd, China) to amplify the sequences of T1 and T2. The reagents and reaction conditions of the PCRs were selected and programmed according to the manufacturer’s instructions. The obtained DNA fragments were sequenced, and then a similarity search using BLAST was performed with the wild-type (WT)sequence.
Identification of transgene-free lines
To identify the transgene-free lines, T1seedlings were examined using PCR with specific primers (Table 1) for the sequences of Cas9 and-resistance gene ().
Evaluation of T1 phenotype
The rice seeds were harvested at maturity and dried in an oven at 38 ºC for two days after manual dehulling. The morphology of the seed endosperms was observed by visual inspection. Potassium iodide (I2-KI, 2%) staining was used to identify the starch types in the transgenic seeds T1generation(Kawagoe et al, 2005). Dried grains were ground into fine flour to pass through a 60 mesh sieve. Then, the AC of the powder was measured using a colorimetric method (Juliano et al, 1981). Scanning electron microscopy (SEM) was also performed to investigate the morphology of the rice starch. The polished rice seeds were cut with the blade of a scalpel, mounted on copper stubs, and coated with gold. All detailed procedures were performed according to Kasem et al (2011). Images were recorded on a Hitachi-S-3000Nscanning electron microscope (Hitachi, Co., Ltd. Japan).

Table 1. Primers used in this study.
Results
Construction of CRISPR/Cas9 system
To delete thegene inrice with the CRISPR/Cas9 technology, two different 20-bp sgRNAs (T1 and T2) at the first exon ofwere selected using the web-based tool CRISPR-P (http://cbi.hzau.edu.cn/cgi-bin/CRISPR), according to the criteria reported by Ma et al (2015). Two target sites were located at 15 and 217 bp downstream of the translation initiation codon (ATG) and were named T1 and T2, respectively (Fig. 1-A). The length between T1 and T2 was 202 bp, according to the coding sequence of. Two sgRNAs were sequentially assembled into pYLCRISPR/Cas9-MH as described by Wang et al (2017c). The T1 and T2 in the plasmid were driven by theandpromoters, respectively. Both Cas9 and, elements of the plasmid, were transcribed from the CaMV 35S promoter (Fig. 1-B).
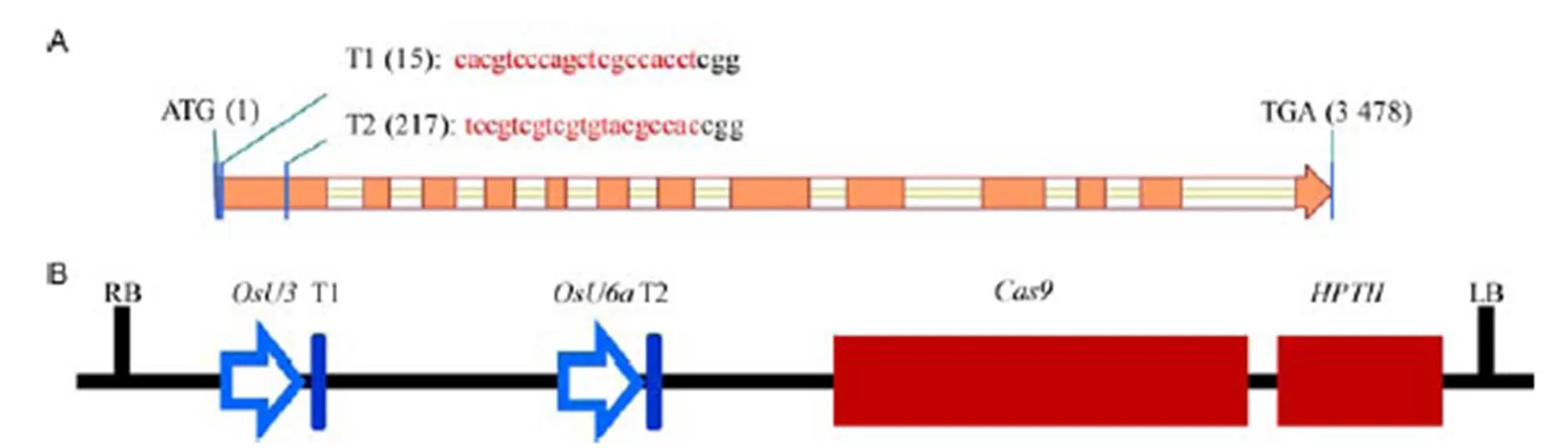
Fig. 1. Schematic diagrams of theand CRISPR/Cas9 vector.
A, Overview of thegene structure and target sites for sgRNAs (T1 and T2). Exons and introns ingene are depicted by orange rectangles and white rectangles, respectively. The target sequences of sgRNAs are shown at the top of the gene structure in red. B, The structure of thegene editing system. T1 and T2 are driven by U3 and U6a promoters, respectively.
RB, Right border; LB, Left border;,CRISPR associated protein 9 gene;,-resistance gene.
Targets editing in T0 plants
The CRISPR/Cas9-construct was transformed into two elite rice inbred lines HD5 and SK118. We obtained 36 and 32 independent T0transgenic plants of HD5 and SK118, respectively. The PCR tests of thegene confirmed that all T0plants were positive transgenic lines (Fig. 2-A). To analyze the mutation genotypes of the T0plants, three mutants of both-HD5 and-SK118 were randomly selected for PCR amplification (Fig. 2-B). The PCR products were sequenced. The genotype differences between the T0mutants and the WT lines are shown in Fig. 2-C. The results showed that the mutagenesis efficiency of the target sites was very high, the deletion fragments were larger than 200 bp in all individuals, the nucleotides between twoprotospacer adjacent motif (PAM) sites of the targets were all deleted by CRISPR/Cas9, and all T0plants were homozygous mutations.
Selection of transgene-free homozygous mutant lines
Because inherited Cas9 could also induce new mutations and lead to unpredicted segregation, distortion and chimerism in later generations, it is important to eliminate Cas9 by segregation in the T1generation and obtain stable mutants (Ishizaki, 2016; Yin et al, 2017). Three-SK118 (-SK118-1,-SK118-2 and-SK118-3) and three-HD5 (-HD5-1,-HD5-2 and-HD5-3) T0mutants were self-pollinated to generate T1plants. Transgene-free lines were analyzed by PCR using specific primers for theandsequences in the T1plants (Table 1). The results were determined using negative PCRs of bothand. The 235-HD5 and 118-SK118T1plants (including selections from all the six lines) were subjected to PCR tests, respectively, and 36-HD5 and 18-SK118transgene-free plants were isolated, respectively (Table 2).
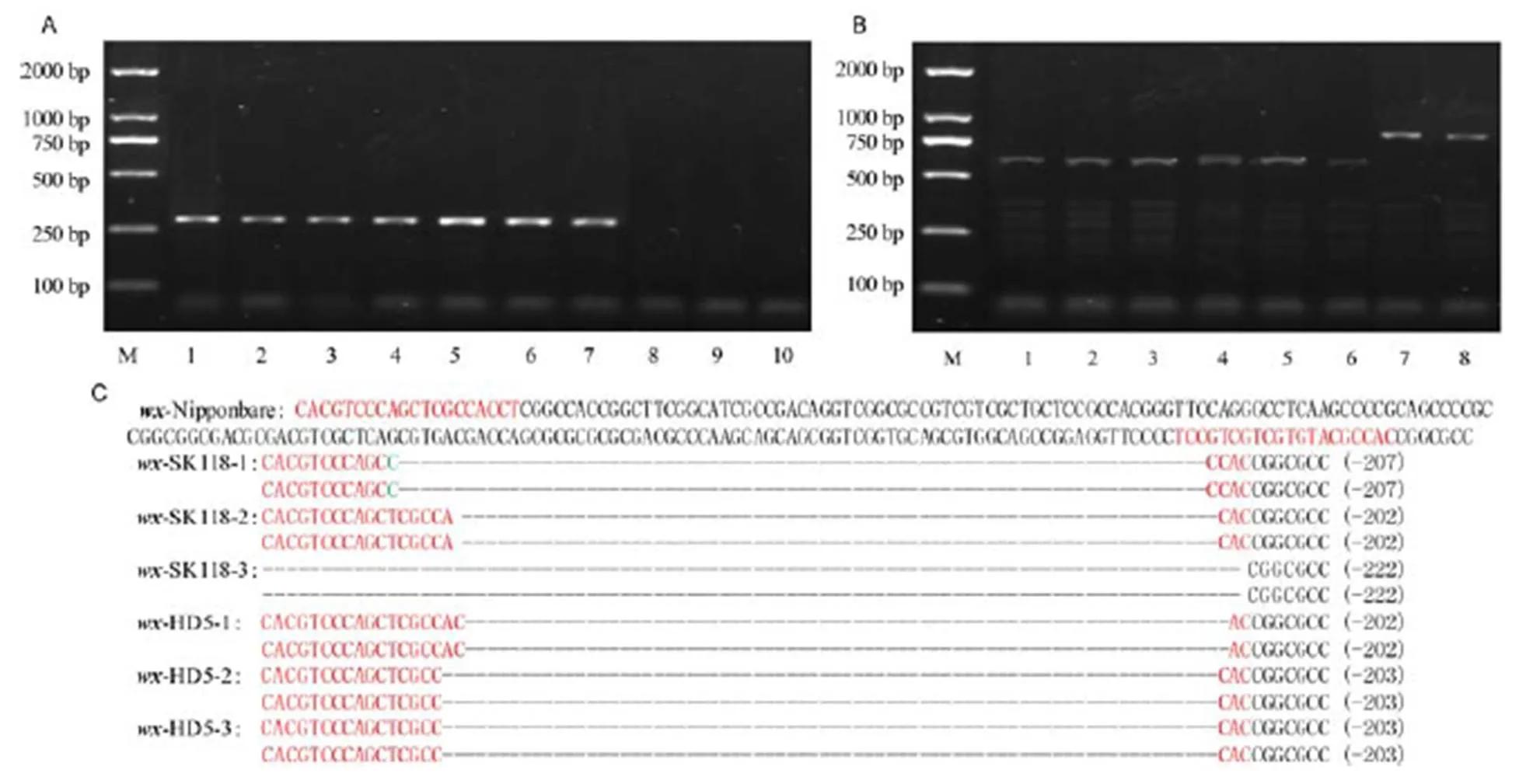
Fig. 2. CRISPR/Cas9-induced mutations in thegene.
A, PCR analysis ofin T0lines (part of the picture). M, DNA marker; 1,-SK118-1; 2,-SK118-2; 3,-SK118-3; 4,-HD5-1; 5,-HD5-2; 6,-HD5-3; 7, Positive control of pYLCRISPR/Cas9-MH-; 8, Negative control of SK118; 9, Negative control of HD5; 10, Negative control of ddH2O. B, Polymerase chain reaction assays ofin T0lines. M, DNA marker; 1,-SK118-1; 2,-SK118-2; 3,-SK118-3; 4,-HD5-1; 5,-HD5-2; 6,-HD5-3; 7, Control of SK118; 8, Control of HD5. C, Sequencing results of T0mutant lines. sgRNA is shown in red letter; Insertion sequence is shown in green letter; Deletion sequence is shown by dashed line.
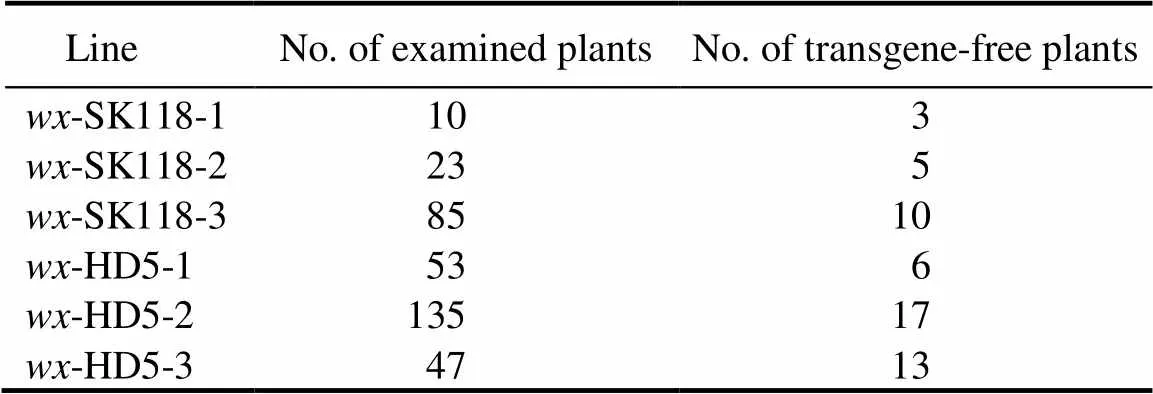
Table 2. Segregation of transgene-free rice in T1 generation.
Amylose content of rice grains
The phenotypes of the mutant grains were compared with those of the WT. As shown in Fig. 3-A, the grains of the mutants were all white and opaque, which showed similar characteristics to those of typical glutinous rice varieties. The starch properties of the mutants were identified by a screen with I2-KI staining, as the seeds with more amylose have higher iodine binding capacity. Non-glutinous rice varieties have granules that are stained with a dark blue color. Seeds with more amylopectin, such as sticky rice, have a lower iodine binding capacity, and therefore, granules are stained with a reddish brown color (Kawagoe et al, 2005). As shown in Fig. 3-B, all the seeds of the mutants were reddish brown, while the WT grains were dark blue. The results revealed that the amylose content in the mutants was dramatically decreased. Further analysis showed that the AC of the mutants was 2.6%–3.2%, while the AC of the WT grains was 15.4%–15.7% (Fig. 3-C). As expected, the dramatic decrease of AC in the mutants was closely associated with the grain appearance and iodine staining results. The-HD5 and-SK118 rice strains were turned into glutinous lines.
Microstructural features
The cross sections of the grains were observed with SEM (Fig. 4). Results in Fig. 4-A revealed that the grain surfaces of the mutants were smoother and flatter than in the wild-types. Radiation from the rectangular columns, as revealed by cross-sections of grains, was reduced in the mutants, especially in-HD5. The fractured surfaces of grains from both the mutants and WTs were all densely packed with starch granules, and all the starch granules displayed large and irregular polygonal shapes with sharp edges (Fig. 4-B).
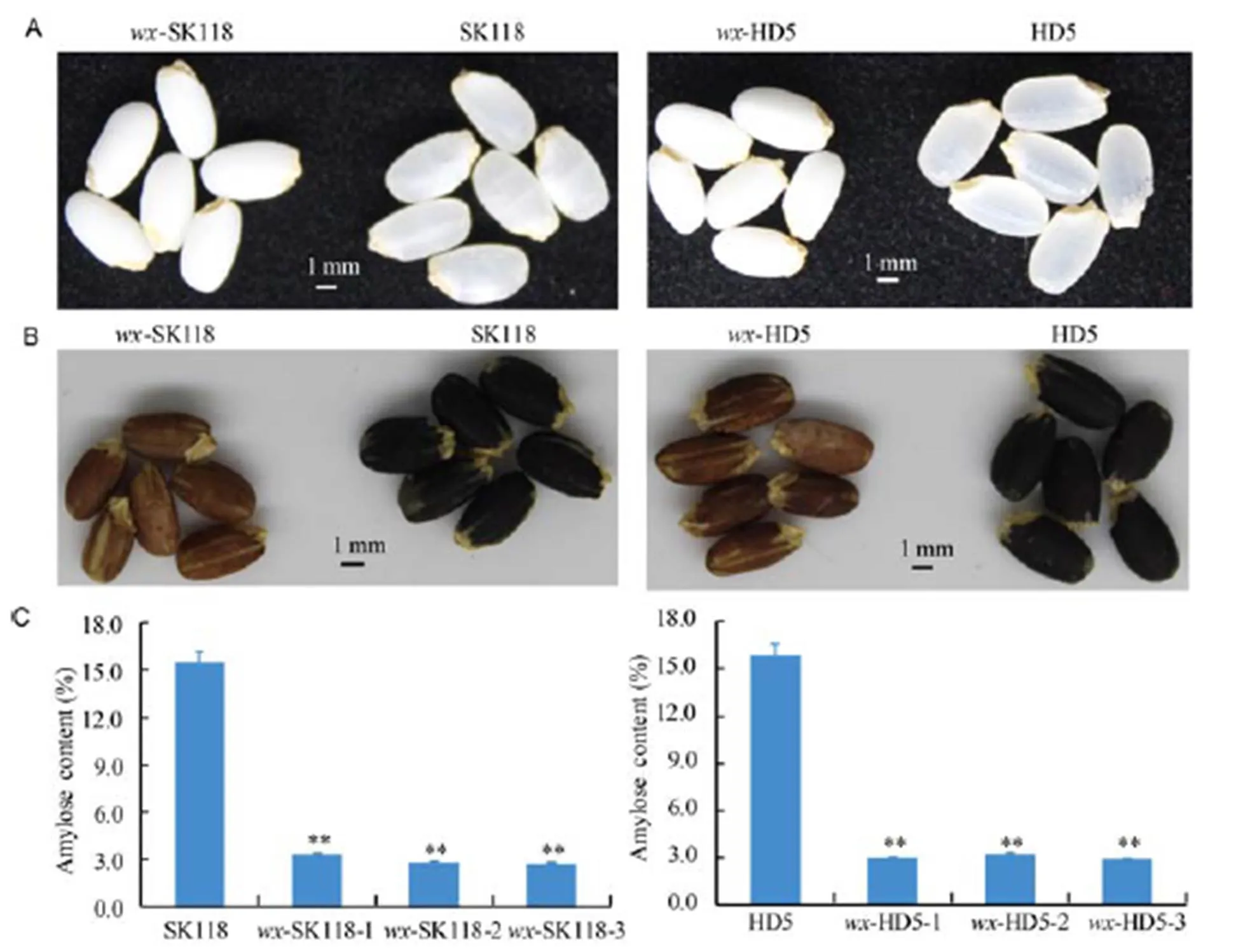
Fig. 3. Phenotype of themutants.
A, Intact polished rice. B, Grains stained with iodine solution. C, Amylose content ofmutants.
Data represent Mean ± SD (= 3). **, Significant at the 0.01 level by the-test.
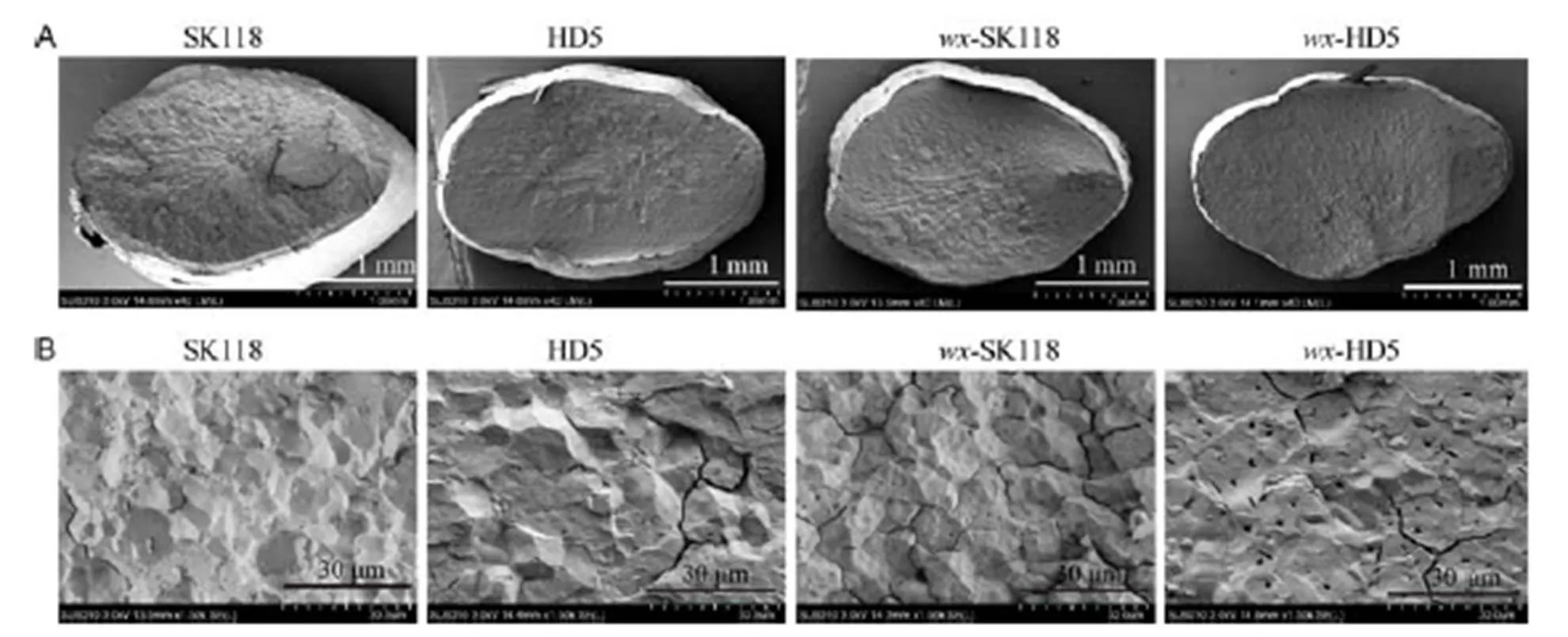

Fig. 4. Cross-sections of endosperm in rice.
DisCussion
The first application of CRISPR/Cas9-based genome editing in rice was presented in 2013 (Cho et al, 2013). Since then, the technology has been widely used to improve agronomic traits and create novel germplasm in rice. Li et al (2016) and Zhou et al (2016) developed photoperiod-controlled and thermo-sensitive male sterile lines by disrupting theandgenes, respectively. Mutants of the,andgenes, which function as negative regulators of grain weight, were generated through CRISPR/Cas9-mediated targeted mutagenesis (Xu et al, 2016). Li et al (2017) designed sgRNAs to target,and, and produced early-maturing rice germplasm, which can improve the ecological adaptability of those rice varieties. Sun et al (2017) created high amylose rice plants through CRISPR/Cas9-mediated target gene editing ofand. The CRISPR/Cas9 technology has a great potential to facilitate plant breeding. Given this, we developed two elite glutinous rice varieties using the CRISPR/Cas9 technology by editing thegene.
We designed two sgRNAs (T1 and T2), targeting the first exon of thegene. The GC contents of T1 and T2 were 70% and 65% respectively, and the secondary structures of sgRNAs had less than 6 bp complete matching. Using these sgRNAs, we observed extremely high editing efficiency in thegene. The design parameters of the sgRNA are in line with the reported critia (Ma et al, 2015), which might be a reason for the high sgRNA editing efficiency in this study. It was very interesting that the genotypes of all mutants generated from six lines were homozygous with more than 200 bp deletions and that the fragment between T1 and T2 was missing. It has been reported that CRISPR/Cas9 induces modifications that are small, presumably because of the blunt-ended double-strand breaks (Zheng et al, 2016). In this study, the CRISPR/Cas9 systerm seemed to preferentially induced full-length deletions, rather than small modifications. When the number of base pairs between the two target sites in Wx was 202 bp, and so that we found that the genetypes of all the six selected T0transgenic plants were about 200 bp-fragment deletion. A similar phenomenon was described by (Wang et al, 2017a), but the distance between the sgRNAs required to induce large deletions should be further explored.
mutation with different amylose content has been reported (Toru and Naoko, 2017). Rice varieties can be classified as waxy (0–2%), very low (3%–9%), low (10%–19%), intermediate (20%–25%) and high (> 25%) according to their AC (Cai et al, 2015). Thegene is the principal gene controlling the AC and can explain almost 90% of the AC variation in some cultivars (Wamnugu et al, 2017). Variousmutations (Wx,Wx,Wx,Wx,Wxand) generate a wide range of variation in AC (Yang et al, 2016; Wang et al, 2018). Studies have reported that different types ofmutations, such as a G/T mutation at the 5′ splice site of intron 1, a C deletion at intron 5, a G/A SNP in thepromoter region, or a T/G SNP at the intron 1-exon 1 junction, would result in amylose content decreased (Wang et al, 2015; Shahid et al, 2016). Ma et al (2015) disrupted thegene by CRISPR/Cas9 system in arice Taichung 65, and the AC in the mutants decreased to 2.6%, which is similar to a natural glutinous rice. Zhang et al (2018) used the CRISPR/Cas9 to target the first exon ofin tworice varieties, Xiushui 134 and Wuyunjing 7, and the results showed that the AC of mutants is reduced without affecting the other desirable agronomic traits. Our results generally agree with those reported in other studies.
In conclusion, we precisely edited thegene using CRISPR/Cas9technology and generated high-quality glutinous rice. This work would provide new materials for traditional breeding and broaden the genetic diversity of glutinous rice.
ACKNOWLEDGEMENTS
This work was supported by the National Key Research and Development Program (Grant No. 2017YFD0100403), the Jiangsu Province Key Research and Development Program (Modern Agriculture) Project (Grant No. BE2017345-2), the Exploratory Project of the Jiangsu Academy of Agricultural Sciences [Grant No. ZX(17)2014], and the Jiangsu Province Natural Science Foundation (Grant No. BK20171326).
Baysal C, Bortesi L, Zhu C, Farre G, Schillberg S, Christou P. 2016. CRISPR/Cas9 activity in the ricegene does not induce off-target effects in the closely related paralog., 36: 108.
Cai J W, Man J M, Huang J, Liu Q Q, Wei W X, Wei C X. 2015. Relationship between structure and functional properties of normal rice starches with different amylose contents., 125: 35–44.
Chen M H, Bergman C, Pinson S, Fjellstrom R. 2008.gene haplotypes: Associations with apparent amylose content and the effect by the environment in an international rice germplasm collection., 47(3): 536–545.
Cho S W, Kim S, Kim J M, Kim JS. 2013. Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease., 31: 230–232.
Fasahat P, Rahman S, Ratnam W. 2014. Genetic controls on starch amylose content in wheat and rice grains., 93(1): 279–292.
Ishizaki T. 2016. CRISPR/Cas9 in rice can induce new mutations in later generations, leading to chimerism and unpredicted segregation of the targeted mutation., 36(12): 165.
Juliano B O, Perez C M, Blakeney A B, Castillo D T, Kongseree N, Laignelet B, Lapis E T, Murty V V S, Paule C M, Webb B D. 1981. International cooperative testing on the amylose content of milled rice., 33(5): 157–162.
Kasem S, Waters D L E, Rice N F, Shapter F M, Henry R J. 2011. The endosperm morphology of rice and its wild relatives as observed by scanning electron microscopy., 4(1): 12–20.
Kawagoe Y, Kubo A, Satoh H, Takaiwa F, Nakamura Y. 2005. Roles of isoamylase and ADP-glucose pyrophosphorylase in starch granule synthesis in rice endosperm., 42: 164–174.
Kiswara G, Lee J H, Hur Y J, Cho J H, Lee J Y, Kim S Y, Sohn Y B, Song Y C, Nam M H, Yun B W. 2014. Genetic analysis and molecular mapping of low amylose gene(t) in rice (L.)., 127(1): 51–57.
Li Q L, Zhang D B, Chen M J, Liang W Q, Wei J J, Qi Y P, Yuan Z. 2016. Development ofphoto-sensitive genic male sterile rice lines by editing carbon starved anther using CRISPR/Cas9., 43(6): 415–419.
Li X F, Zhou W J, Ren Y K, Tian X J, Lv T X, Wang Z Y, Fang J, Chu C C, Yang J, Bu Q Y. 2017. High-efficiency breeding of early-maturing rice cultivars via CRISPR/Cas9-mediated genome editing., 44(3): 175–178.
Lu G, Moriyama E N. 2004. Vector NTI, a balanced all-in-one sequence analysis suite., 5(4): 378–388.
Ma X L, Zhang Q Y, Zhu Q L, Liu W, Chen Y, Qiu R, Wang B, Yang Z F, Li H Y, Lin Y R, Xie Y Y, Shen R X, Chen S F, Wang Z, Chen Y L, Guo J X, Chen L T, Zhao X C, Dong Z C, Liu Y G. 2015. A robust CRISPR/Cas9 system for convenient, high- efficiency multiplex genome editing in monocot and dicot plants., 8(8): 1274–1284.
Mikami M, Toki S, Endo M. 2015. Comparison of CRISPR/Cas9 expression constructs for efficient targeted mutagenesis in rice., 88(6): 561–572.
Murray M G, Thompson W F. 1980. Rapid isolation of high molecular weight plant DNA., 8(19): 4321–4325.
Nieves-cordones M, Mohamed S, Tanoi K, Kobayashi N I, Takagi K, Vernet A, Guiderdoni E, Périn C, Sentenac H, Véry A A. 2017. Production of low-Cs(+) rice plants by inactivation of the K(+) transporter OsHAK1 with the CRISPR-Cas system., 92(1): 43–56.
Peng B, Sun Y F, Pang R H, Li H L, Song X H, Yuan H Y, Zhang S H, Zhou Q Y, Li Q R, Li D, Song S Z. 2016. Study on chalkiness character and endosperm structure of rice grain in differentvarieties., 28(11): 1803–1811. (in Chinese with English abstract)
Shahid S, Begum R, Razzaque S, Jesmin, Seraj Z I. 2016. Variability in amylose content of Bangladeshi rice cultivars due to unique SNPs inallele., 71: 1–9.
Shen L, Hua Y F, Fu Y P, Jian L, Liu Q, Jiao X Z, Xin G W, Wang J J, Wang X C, Yan C J, Wang K J. 2017. Rapid generation of genetic diversity by multiplex CRISPR/Cas9 genome editing in rice., 60(5): 506–515.
Sun Y W, Jiao G A, Liu Z P, Zhang X, Li J Y, Guo X P, Du W M, Du J L, Francis F, Zhao Y D, Xia L Q. 2017. Generation of high-amylose rice through CRISPR/Cas9-mediated targeted mutagenesis of starch branching enzymes., 8: 298.
Tang L, Mao B G, Li Y K, Lv Q M, Zhang L P, Chen C Y, He H J, Wang W P, Zeng X F, Shao Y, Pan Y L, Hu Y Y, Peng Y, Fu X Q, Li H Q, Xia S T, Zhao B R. 2017. Knockout ofusing the CRISPR/Cas9 system produces low Cd-accumulatingrice without compromising yield., 7(1): 14438.
Toru T, Naoko F. 2017. Thermal and rheological characteristics of mutant rice starches with widespread variation of amylose content and amylopectin structure., 62: 83–93.
Wamnugu P, Ndjiondjop M N, Furtado A, Henry R. 2017. Sequencing of bulks of segregants allows dissection of genetic control of amylose content in rice., 16: 100–110.
Wang B K, Zhang H, Hong R K, Zhang J W, Yang R, Luo Q, Zeng Q C. 2018.gene editing via CRISPR/Cas9 system in rice., 32(1): 35–42. (in Chinese with English abstract)
Wang F J, Wang C L, Liu P Q, Lei C L, Hao W, Gao Y, Liu Y G, Zhao K J. 2016. Enhanced rice blast resistance by CRISPR/Cas9-targeted mutagenesis of the ERF transcription factor gene., 11(4): e0154027.
Wang F Q, Fan F J, Li W Q, Zhu J Y, Wang J, Zhong W G, Yang J. 2016. Knock out efficiency analysis ofgene using CRISPR/Cas9 in rice., 30(5): 469–478. (in Chinese with English abstract)
Wang K, Hasjim J, Wu A C, Li E, Henry R J, Gilbert R G. 2015. Roles ofandin determining amylose fine structure., 127: 264–274.
Wang M G, Mao Y F, Lu Y M, Tao X P, Zhu J K. 2017a. Multiplex gene editing in rice using the CRISPR-Cpf1 system., 10(7): 1011–1013.
Wang Q X, Zhao H, Jiang J P, Xu J Y, Xie W B, Fu X K, Liu C, He Y Q, Wang G W. 2017b. Genetic architecture of natural variation in rice nonphotochemical quenching capacity revealed by genome-wide association study., 8: 1773.
Wang Y, Geng L Z, Yuan M L, Wei J, Jin C, Li M, Yu K, Zhang Y, Jin H B, Wang E, Chai Z J, Fu X D, Li X G. 2017c. Deletion of a target gene inrice via CRISPR/Cas9., 36(8): 1333–1343.
Wu Y P, Pu C H, Lin H Y, Huang H Y, Huang Y C, Hong C Y, Chang M C, Lin Y R. 2015. Three novel alleles of() confer dull grains with low amylose content in rice., 233: 44–52.
Xie Y Y, Niu B X, Long Y M, Li G S, Tang J T, Zhang Y L, Ren D, Liu Y G, Chen L T. 2017. Suppression or knockout of/overcomes the-mediated hybrid male sterility in rice., 59(9): 669–679.
Xu R F, Yang Y C, Qin R Y, Li H, Qiu C H, Li L, Wei P C, Yang J B. 2016. Rapid improvement of grain weight via highly efficient CRISPR/Cas9-mediated multiplex genome editing in rice., 43(8): 529–532.
Yang X H, Nong B X, Xia X Z, Zhang Z Q, Zeng Y, Liu K Q, Deng G F, Li D T. 2016. Rapid identification of a new gene influencing low amylose content in rice landraces (L.) using genome-wide association study with specific-locus amplified fragment sequencing., 60(6): 465–472.
Yin X J, Biswal A K, Dionora J, Perdigon K M, Balahadia C P, Mazumdar S, Chater C, Lin H C, Coe R A, Kretzschmar T, Guick J E, Quick P W, Bandyopadhyay A. 2017. CRISPR-Cas9 and CRISPR-Cpf1 mediated targeting of a stomatal developmental genein rice., 36(5): 745–757.
Zhang H, Xu H, Feng M J, Zhu Y. 2018. Suppression ofin rice endosperm stabilizes amylose content under high temperature stress., 16(1): 18–26.
Zhang J S, Zhang H, Botella J R, Zhu J K. 2018. Generation of new glutinous rice by CRISPR/Cas9-targeted mutagenesis of thegene in elite rice varieties., 60(5): 369–375.
Zheng X L, Yang S X, Zhang D W, Zhong Z H, Tang X, Deng K J, Zhou J P, Qi Y P, Zhang Y. 2016. Effective screen of CRISPR/Cas9-induced mutants in rice by single-strand conformation polymorphism., 35(7): 1545–1554.
Zhou H, He M, Li J, Chen L, Huang Z F, Zheng S Y, Zhu L Y, Ni E, Jiang D G, Zhao B R, Zhuang C X. 2016. Development of commercial thermo-sensitive genic male sterile rice accelerates hybrid rice breeding using the CRISPR/Cas9-mediatedediting system., 6: 37395.
(Managing Editor: Wang Caihong)
Copyright © 2019, China National Rice Research Institute. Hosting by Elsevier B V
This is an open access article under the CC BY-NC-ND license (http://creativecommons.org/licenses/by-nc-nd/4.0/)
Peer review under responsibility of China National Rice Research Institute
http://dx.doi.org/10.1016/j.rsci.2018.04.007
26 February 2018;
27 April 2018
Yang Jie (yangjie168@aliyun.com)
杂志排行
Rice Science的其它文章
- Editing of Rice Isoamylase Gene ISA1 Provides Insights into Its Function in Starch Formation
- Characterization and Evaluation of OsLCT1 and OsNramp5 Mutants Generated Through CRISPR/Cas9-Mediated Mutagenesis for Breeding Low Cd Rice
- Targeted Mutagenesis of NAC Transcription Factor Gene, OsNAC041, Leading to Salt Sensitivity in Rice
- CRISPR/Cas9-Mediated Adenine Base Editing in Rice Genome
- Rapid Creation of New Photoperiod-/Thermo-Sensitive Genic Male-Sterile Rice Materials by CRISPR/Cas9 System
- Development and Application of CRISPR/Cas System in Rice
