Isolation and Identification of Canine Enterococcus faecalis
2019-09-10YanfeiLIULuLUXimoZHANGJiandeYANG
Yanfei LIU Lu LU Ximo ZHANG Jiande YANG

Abstract With the development of the urban pet industry, suppurative diseases of companion animals are increasing day by day. To explore its associated pathogens, this study was conducted on etilolgical identification from a case of a canine uterus with pus. Bacteriological detection methods were used to isolate and purify the collected disease materials, such as smear staining, microscopic examination and biochemical identification. Then, 16S rRNA molecular sequence analysis was used to identify the isolate. It was found that the morphological features of the isolated strain and the biochemical results were consistent with Enterococcus faecalis. The 16S rRNA of the isolate was also as high as 99% homologous with the reference stains. Drug susceptibility test showed that the isolate was resistant to tetracycline, penicillin G, minocycline, erythromycin, ciprofloxacin, norfloxacin, vancomycin, levofloxacin, clindamycin, polymyxin B, chloramphenicol, compound sulfamethoxazole, and clarithromycin, and only susceptible to oxacillin and nitrofurantoin. These results provided reference for the treatment of this type of multidrug resistant enterococcal infection.
Key words Enterococcus faecalis; Biochemical identification; Evolutionary tree analysis; Drug sensitive test
Enterococcus faecalis, also known as Streptococcus faecalis, is positive by Gram staining and is one of the major flora in the intestine tract of humans and animals[1]. E. faecalis can grow and survive in different environments, and often settles in the intestine, mouth and genital tract of animals. It can produce natural antibiotics, which is beneficial to the health of the body[2]. In the past, E. faecalis was considered to be a symbiotic bacterium of humans and animals. However, in the past decade, E. faecalis has gradually become the second largest pathogen of Gram positive bacteria in humans, second only to Staphylococcus[3]. E. faecalis can frequently cause widespread infection in humans, and the number of cases of infection and death in hospitals is gradually increasing, especially in patients with low immunity or patients with critical underlying diseases[4]. Most of these infections occur in urinary tracts, blood circulation, endocardial abdomens, bile ducts, burns, and the like. E. faecalis can also infect the central nervous system, lungs, soft tissues, paranasal sinuses, eyes and periodontal tissues, but these tissues are less frequently infected[5]. In many European and American countries, E. faecalis infection is a serious threat to human public health. In China, pathogenic E. faecalis has also been isolated from animals such as pigs and sheep[6]. In recent years, reports of the incidence of E. faecalis in animals and the death of animals caused thereby have been increasing , often causing diseases such as boar orchitis, chicken sepsis, bovine mastitis, calf or lamb endocarditis and canine urinary tract infections[7]. The large scale use of antibiotics has led to the evolution of many antibiotic resistant mechanisms in E. faecalis and the production of corresponding resistance mechanisms[8].
In this study, the pus in the uterus of the affected dog was collected, and a bacterial strain was obtained by isolation and culture, and named E f lu, which was subjected to morphological observation, biochemical test, drug susceptibility test, and amplification and sequencing analysis of 16S rRNA.
Materials and Methods
Tissue samples
The sick dog was a 10 year old female German shepherd dog sent to the Animal Hospital Affiliated to Tianjin Agricultural University. The symptoms were yellow vaginal discharge, unpleasant smell and slightly enlarged abdominal circumference. The pus uterus was removed surgically, and the pus was collected in a sterile environment for pathogen isolation.
Main reagents
Media and microbiological identification tube
Common broth medium, nutrient agar medium, Columbia agar medium and MacConkey agar medium were all prepared according to the conventional method[9]. The Streptococcus identification tubes were purchased from Hangzhou Binhe Microorganism Reagent Co., Ltd.
Antimicrobial susceptibility disk
The antimicrobial susceptibility disks were purchased from Hangzhou Binhe Microorganism Reagent Co., Ltd.
Methods
Isolation and culture of bacteria
The pus was aseptically inoculated on the medium, and then the dominant colonies were selected for pure culture. They were streaked on common nutrient agar plates and Columbia agar plates, and cultured in a 37 ℃ incubator for 24 h, and then stored in a refrigerator at 4 ℃ for later use.
Staining microscopy
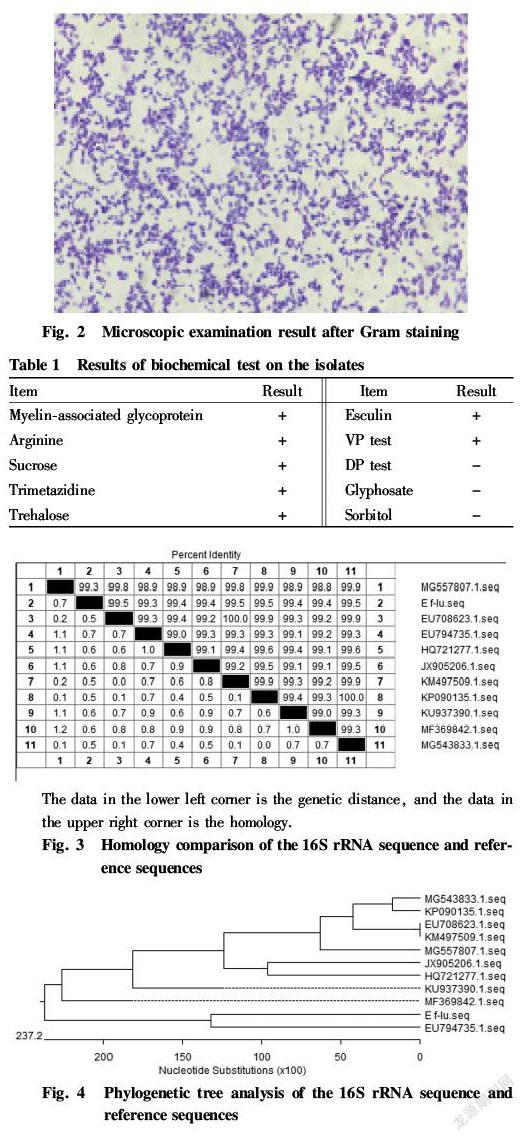
Individual colonies were picked for smear preparation and Gram staining, and the morphology, arrangement and staining characteristics of the bacteria were observed under oil lens. The staining was carried out according to the requirements of reference[10].
Identification of biochemical characteristics
The biochemical test was referred to "Manual for Systematic Identification of Common Bacteria", and the pure culture of the isolated bacteria was inoculated into the biochemical identification tubes of arginine, sucrose, esculin, VP test, etc., and cultured at 37 ℃ for 24 h. The biochemical reaction was observed, and the results were recorded.
16S rRNA homology analysis and phylogenetic tree construction of the isolate
The obtained isolate was sent to Sangon Biotech (Shanghai) Co., Ltd. to sequence the 16S rRNA gene sequences, and the obtained gene sequences were subjected to Blast similarity search in the NCBI database to obtain the sequences closest to the 16S rRNA sequence of the isolate. Multi sequence homology alignment and analysis and the construction of the phylogenetic trees were performed using DNAstar software.
Susceptibility test
The plate surface dispersion method was applied. 100 μl (109 CFU) of the broth culture of each isolate was evenly spread on the prepared nutrient agar plate, and the drug susceptibility disk was attached to the nutrient agar medium. The plates with the susceptibility disk were cultured under an inverted condition at 37 ℃ for 24 h in an incubator. The inhibitory zone diameters were observed and recorded in the drug susceptibility test, for the analysis of drug susceptibility.
Results
Isolation and culture characteristics, bacterial morphology and dyeing characteristics
The isolated bacteria formed small rounded milky white opaque colonies with smooth surface and neatly margin in nutrient agar, as shown in Fig. 1. It showed slightly larger colonies on Columbia agar, but did not grow on MacConkey medium. The bacteria were subjected to smear staining and microscopic examination, which showed that the isolate was Gram positive cocci, mostly in a double spherical arrangement, and a few were single, as shown in Fig. 2.
Results of biochemical test
The isolate was tested to be positive for myelin associated glycoprotein, arginine, sucrose, trimetazidine, trehalose, esculin and VP test, and negative for DP test, glyphosate and sorbitol. With reference to the physiological and biochemical characteristics of the standard strain, the isolate was consistent with enterococcus, and was therefore initially identified as enterococcus. The details are shown in Table 1.
Analysis of 16S rRNA homology results
This sequence was subjected to Blast alignment on the NCBI website and found to be up to 100% similar to E. faecalis. The 16S rRNA gene sequence of the isolate was analyzed by DNAstar software with 16S rRNA in the database. The results showed that the 16S rRNA sequence of the isolate shared high homology with E. faecalis, and the homology was 99%, as shown in Fig. 3.
Analysis results of 16S rRNA gene phylogenetic tree
A phylogenetic tree was constructed with the 16S rRNA sequence of the isolate and 10 sequences in the database by MegAlign in DNAstar software. The results are shown in Fig. 4. The isolate E f lu had the closest affinity to EU794735.1 and MF369842.1 in the database, and was relatively distantly related to MG543833.1 and KP090135.1. It shared the highest homology with EU708623.1, KM497509.1, KP090135.1 and MG543833.1, all at 99.5%.
Drug susceptibility test
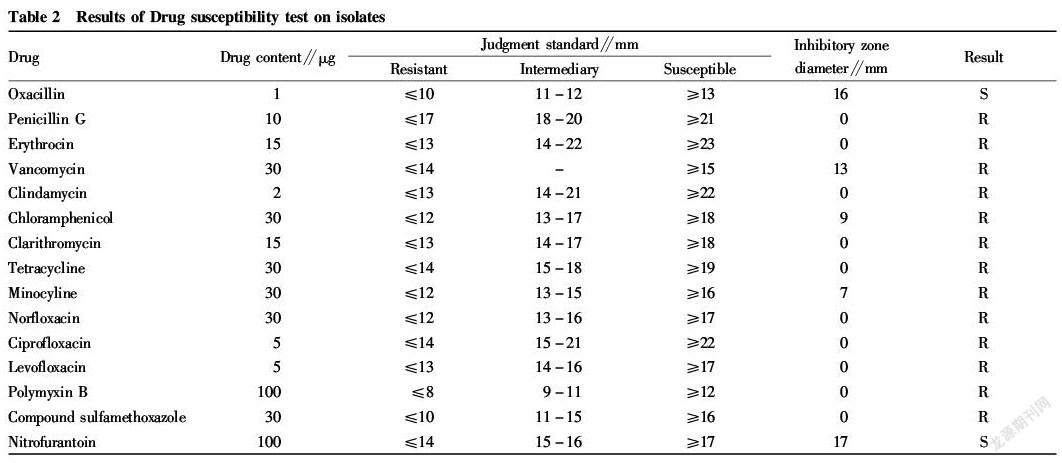
The resistance of the isolate to vancomycin, minocycline, chloramphenicol, oxacillin, clindamycin and other antibiotics was determined, indicating that the isolate is substantially resistant to all the drugs tested, only sensitive to oxacillin and nitrofurantoin. The results are shown in Table 2 in details.
Discussion
E. faecalis can cause diseases such as genitourinary tract infection, bacteremia, sepsis, traumatic infection, endocarditis and skin soft tissue infections[5]. In 2010-2014, Jiang et al.[11] isolated from the clinical specimens from the hospital where they worked, 1 220 E. faecalis isolates, including 675 E. faecalis isolates (55.3%) and 445 E. faecium isolates (36.5%). Although E. faecalis predominates in clinical infections, it is rare in clinical canine pyogenic diseases. In 2013, Shi[12] conducted pathogen identification of 43 cases of canine pyometra, and found that the main isolates were Gram negative bacilli, most of which were members of Enterobacteriaceae, and a small part were Pseudomonas aeruginosa in Pseudomonas; and Gram positive cocci was also isolated, including E. faecalis and Staphylococcus, but the proportion was only 15% of the total pyometra.
In this study, the E. faecalis isolated from the pus of the canine pyometra was tested for drug susceptibility test. It was tested to be resistant to vancomycin, minocycline, clarithromycin, chloramphenicol, etc., only susceptible to oxacillin and nitrofurantoin, indicating that the isolate is a multi drug resistant strain. Even the intractable vancomycin has been broken through by E. faecalis, which has attracted great attention from China Antimicrobial Surveillance Network (CHINET)[13].
Due to the prevalence of E. faecalis in pigs, sheep, cows, poultry farms, hospitals and animal clinics, the long term use of antibiotics has led to an increase in the resistance to E. faecalis, making the treatment of E. faecalis infection more and more difficult. Currently, vaccines for the prevention of E. faecalis disease are not yet widespread and are still in the laboratory research stage[14]. Multi drug resistant bacteria can spread among humans, animals and the environment, making the problem of resistance in bacteria more serious and posing a serious threat to human public health[15]. Therefore, the treatment of E. faecalis infection should not only clarify its main pathogenic mechanism, but also strengthen the rational and standardized use of human and animal antibiotics.
References
[1] WANG YB, CHEN LY, HU H, et al. Development of a multiplex PCR assay for detection of swine originated E. faecalis and E. faecium isolates[J]. Chinese Journal of Veterinary Science, 2011, 31(8): 1123-1127. (in Chinese)
[2] MALIK RK, MONTECALVO MA, REALE MR, et al. Epidemiology and control of vancomycin resistant enterococci in a regional neonatal intensive care unit[J]. Pediatr Infect Dis J, 1999, 18(4): 352.
[3] TOLEDO ARANA A, VALLE J, SOLANO C, et al. The enterococcal surface protein, Esp, is involved in Enterococcus faecalis biofilm formation[J]. Appl Environ Microbiol, 2001, 67(10): 4538-4545.
[4] CRETI R, IMPERI M, BERTUCCINI L, et al. Surver for virulence determinants among Enterococcus faecalis isolated from different sources[J]. J Med Microbiol, 2004, 53: 13-20.
[5] LIU YF, YANG JD, ZHAO RL. Research progress of animal diseases caused by Enterococcus faecalis[J]. Journal of Domestic Animal Ecology, 2013, 34(11): 75-77. (in Chinese)
[6] WANG YB, CHEN LY, ZHANG HY, et al. Identification and determination of virulence gene for different isolates of E. faecalis originated from infective piglet[J]. Chinese Journal of Veterinary Science, 2010, 30(5): 615-619. (in Chinese)
[7] ANA RF, ANA PT, CARLA N, et al. Global spread of the hylemf colonization virulence gene plasmids of the Enterococcus farcium CC17 polyclonal sub cluster[J]. Chemother, 2010, 54(6): 2660-2665.
[8] WANG SL, LI YF, LIU QJ, et al. Drug resistance of swine Enterococcus faecalis in hunan province[J]. Chinese Journal of Animal Infectious Diseases, 2015, 23(2): 41-46. (in Chinese)
[9] HU GX, LIAO M, PENG YY, et al. Veterinary microbiology experiment course[M]. Beijing: China Agriculture Press, 2006. (in Chinese)
[10] CUI BA, NIU ZX, REN J, et al. Animal microbiology[M]. Beijing: China Agriculture Press, 2005. (in Chinese)
[11] JIANG DY, LIN XF, WANG BY, et al. Clinical distribution and drug resistance analysis of hospital infection en terococci[J]. China Modern Doctor, 2015, 53(35): 99-102. (in Chinese)
[12] SHI CY. Isolation of bacteria from 43 cases of canine pyometra and the histopathology and immunohistochemical changes of uterine wall[D]. Yangzhou: Yangzhou University, 2016. (in Chinese)
[13] HU FP, ZHU DM, WANG X. 2011 CHINET surveillance of bacterial resistance in China[J]. Chinese Journal of Infection and Chemotherapy, 2014, 14(5): 365-374. (in Chinese)
[14] LYU N, GAO Y. Advances in research on pathogenic factor of Enterococcus faecalis[J]. Journal of Inner Mongolia University for Nationalities: Natural Sciences, 2015, 30(4): 323-326. (in Chinese)
[15] WANG J, WANG XH, XU H. Antimicrobial resistance and dissemination of multidrug resistant organisms A review[J]. Acta microbiologica Sinica, 2016, 56(11): 1671-1679. (in Chinese)
杂志排行
农业生物技术(英文版)的其它文章
- Study on Chemical Composition of the Ethyl Acetate Extract of Pratia
- Effects of Different Selenium Fertilizer Types on Selenium Content and Quality of “Lingfeng” Grapes
- Protoplast Culture and Its Application in Fruit Breeding
- Indexes of Tree Structure of Cylindrical Pear Orchards at the Sapling Stage
- Carbon Storage and Distribution of the Mature Pinus massoniana Plantation in Northwest Guangxi
- Pollution and Quality Control of Mycotoxins in Foods and Feeds
