Long Non-coding RNAs in the Cytoplasm
2016-06-02FarooqRashidAbdullahShahGeShan
Farooq Rashid,Abdullah Shah,Ge Shan
Long Non-coding RNAs in the Cytoplasm
Farooq Rashid#,a,Abdullah Shah#,b,Ge Shan*,c
CAS Key Laboratory of Innate Immunity and Chronic Disease,CAS Center for Excellence in Molecular Cell Science,School of Life Sciences,University of Science and Technology of China,Hefei 230027,China
Received 8 January 2016;revised 3 February 2016;accepted 2 March 2016 Available online 6 May 2016
Handled by Er-Wei Song
KEYWORDS
lncRNA;
mRNA stability;
mRNA translation;
ceRNA;
MicroRNA
Abstract An enormous amount of long non-coding RNAs(lncRNAs)transcribed from eukaryotic genome are important regulators in different aspects of cellular events. Cytoplasm is the residence and the site of action for many lncRNAs. The cytoplasmic lncRNAs play indispensable roles with multiple molecular mechanisms in animal and human cells. In this review,we mainly talk about functions and the underlying mechanisms of lncRNAs in the cytoplasm. We highlight relatively well-studied examples of cytoplasmic lncRNAs for their roles in modulating mRNA stability,regulating mRNA translation,serving as competing endogenous RNAs,functioning as precursors of microRNAs,and mediating protein modifications. We also elaborate the perspectives of cytoplasmic lncRNA studies.
E-mail: shange@ustc.edu.cn(Shan G).
#Equal contribution.
aORCID: 0000-0003-1017-7769.
bORCID: 0000-0003-0130-9133.
cORCID: 0000-0002-3561-2088.
Peer review under responsibility of Beijing Institute of Genomics,Chinese Academy of Sciences and Genetics Society of China.
http://dx.doi.org/10.1016/j.gpb.2016.03.005
1672-0229Ⓒ2016 The Authors. Production and hosting by Elsevier B.V. on behalf of Beijing Institute of Genomics,Chinese Academy of Sciences and Genetics Society of China.
This is an open access article under the CC BY license(http://creativecommons.org/licenses/by/4.0/).
Introduction
Mammalian genome is pervasively transcribed into many different complex families of RNA. However,less than 2% of mammalian genome is transcribed into mRNA to encode proteins,whereas a major portion of the genome is transcribed into interweaved and overlapping transcripts that include thousands of non-coding RNA(ncRNA)transcripts[1,2]. ncRNAs more than 200 nucleotides in length are called long ncRNAs(lncRNAs),which are often transcribed by RNA polymerase II[3,4]. These lncRNAs are usually devoid of open reading frames(ORFs),with or without the 3'polyadenylation [5-8]. Interestingly,expression of lncRNA is more tissuespecific than that of mRNA[9].
In the last several years,a large number of nuclear lncRNAs have been discovered. These lncRNAs play diverse roles in the nucleus through various mechanisms[10]. For example,nuclear lncRNAs control the epigenetic state of particular genes[11],participate in transcriptional regulation[12],get involved in alternative splicing and constitute subnuclear compartments[13,14].
Although for most if not all of the lncRNAs,nucleus is the place of biogenesis and processing,cytoplasm is the final residence and site of action for some lncRNAs. Biogenesis of lncRNAs is quite complicated and share many features of protein-coding RNAs. Within the nucleus,they occupy the chromatin fraction. 17%of lncRNAs vs. 15%of mRNAs are enriched in the nucleus,whereas 4%vs. 26%,respectively,are enriched in the cytoplasm[6]. Many lncRNA-mediated mechanisms of gene regulation have been identified in thecytoplasm[8,15,16]. In the last decade or so,thousands of cytoplasmic lncRNAs have been discovered,indicating their importance for multiple cellular activities. In this review we highlight the functions and underlying mechanisms of some important cytoplasmic lncRNAs that are responsible for posttranscriptional regulations such as on mRNA stability and translational control.
Modulation of mRNA stability
In the cytoplasm,several lncRNAs target mRNA transcripts and modulate mRNA stability. Some lncRNAs such as half-STAU1-binding site RNAs(1/2-sbsRNAs)and growth arrested DNA-damage inducible gene 7(gadd7)decrease the stability of mRNA,while others such as antisense transcript for β-secretase 1(BACE1-AS)and the terminal differentiation-induced ncRNA(TINCR)increase mRNA stability.
1/2-sbsRNAs
mRNAs can be degraded via staufen 1(STAU1)-mediated mRNA decay(SMD),when their 3'untranslated region(3'UTR)binds to STAU1[17]. STAU1 is a double-stranded RNA(dsRNA)-binding protein,which binds within 3'UTR of translationally-active mRNA[14,18]. STAU1 binds to a complex structure of 19-bp stem with 100 nt apex within the mRNA encoding ADP ribosylation factor 1(ARF1)[18]. This stem region is conserved in 3'UTRs of ARF1 mRNA of mouse and rat[6]. However,such stem structures were not identified in other STAU1 targets[18]. STAU1-binding sites can be formed by imperfect base-pairing between an Alu element in the 3'UTR of an SMD target and another Alu element in a cytoplasmic,polyadenylated lncRNA[18]. These lncRNAs transactivate the binding of STAU1 to mRNA as only STAU1 could be immunoprecipitated with lncRNAs called 1/2-sbsRNAs,thus unveiling a pivotal strategy of recruiting proteins to mRNAs and mediating the mRNA decay(Figure 1A). However,not all mRNAs containing Alu element in their 3'UTR are targeted for SMD,despite the presence of complementary 1/2-sbsRNAs that target other mRNA for SMD [17]. One of the 378 identified 1/2-sbsRNAs in humans,1/2-sbsRNA1 contains a single Alu element that base pairs with the Alu element in the 3'UTR of plasminogen activator inhibitor type 1(SERPINE1)and FLJ21870. 1/2-sbsRNA1 is present in the cytoplasm but absent in the nucleus of HeLa cells[18]and only STAU1 can be immunoprecipitated with 1/2-sbsRNA1. Two isoforms of 1/2-sbsRNA1,including 1/2- sbsRNA1(S)(short form)and 1/2-sbsRNA1(L)(long form),have been reported. Both isoforms contain the Alu element and 3’UTR with poly(A)tail,although they differ at the 5'end. Knocking down 1/2-sbsRNA1(S)increased the level of SERPINE1 and FLJ21870 mRNAs by 2-4.5-folds above normal. Other 1/2-sbsRNA members such as 1/2-sbsRNA2,1/2-sbsRNA3,and 1/2-sbsRNA4 are largely cytoplasmic and polyadenylated as well,containing a single Alu element. Knocking down these 1/2-sbsRNAs led to upregulation of their mRNA targets[17]. Functional studies showed that 1/2-sbsRNA1 contributed to the reduced cell migration by targeting SERPINE1 and RAB11-family-interacing protein 1 (RAB11FIP1)mRNAs for SMD as confirmed by scarp injury repair assay[17].
gadd7
gadd7 is a 754-nt polyadenylated lncRNA isolated from Chinese hamster ovary(CHO)cells[15,16]. Expression of gadd7 is induced by several types of DNA damage and growth arrest signals[19,20],and gadd7 plays a pivotal role in regulating G1/S checkpoint post DNA damage. gadd7 also regulates lipid-induced oxidative and endoplasmic reticulum(ER)stress [21]. lncRNAs are known to bind to and regulate the functions of proteins. One such example is the binding of gadd7 with TAR DNA-binding protein(TDP-43),and this interaction is strengthened upon UV exposure[22-24]. TDP-43 is a member of the heterogeneous nuclear ribonucleoprotein(hnRNP)family. HnRNP family members are RNA/DNA binding proteins involved in transcription,splicing,mRNA transport,and mRNA stability[25,26]. TDP-43 is known to repress the expression of cyclin-dependent kinase 6(Cdk6)mRNA in Hela cells,which is important for G1-phase progression[27,28]. Nonetheless,Cdk6 expression is found to be activated by TDP-43 in CHO cells[24]. UV-induced gadd7 directly interacts with TDP-43,thus leading to the decreased interaction between TDP-43 and Cdk6 mRNA. This results in Cdk6 mRNA degradation,and finally inhibition of cell cycle progression[24]. gadd7 is not highly conserved at the nucleotide level[29,30]. Since the structure or the functional motif of lncRNAs may be more important,and thus would be more conserved than their nucleotide sequence[9],it remains possible to identify a functional gadd7 ortholog in humans. This may be important for unveiling the pathogenesis of diseases such as frontotemporal lobar degeneration(FTLD)and amyotrophic lateral sclerosis(ALS),as dominant mutations in TDP-43 are causative of these two important neurodegenerative diseases[24,31,32].
BACE1-AS
Expression of the conserved non-coding BACE1-AS increases BACE1 mRNA stability when HEK-SW cells are exposed to cellular stressors like amyloid-β1-42(Aβ1-42)[33]. BACE1-AS renders BACE1 mRNA stability by masking the binding site of miR-485-5p(Figure 1A). BACE1-AS and miR-485-5p compete for binding in the sixth exon of BACE1 mRNA. The sense-antisense RNA duplex between BACE1 and BACE1-AS in the cytoplasm potentially perturb the interaction between miR-485-5p and BACE1 mRNA,which to some extent,explains the mRNA stabilization by BACE1-AS transcript[34].
TINCR
The TINCR gene resides on chromosome 19 in humans and encodes a predominantly cytoplasmic 3.7-kb lncRNA. TINCR regulates human epidermal differentiation by post transcriptional mechanism[35]. Previously found as an uncharacterized lncRNA,TINCR is now believed to be the most highlyinduced lncRNA during epidermal differentiation[35,36]. TINCR binds to mRNA through a 25-nt‘TINCR box’motif,which is robustly enriched in the interacted mRNAs. TINCR RNA has a strong affinity for STAU1 protein[17,35,37,38]. TINCR-STAU1 complex mediates the stabilization of differentiation-related mRNAs,such as KRT80 encodingkeratin 80 in an ultraviolet protection factor 1/2(UPF1/2)-independent manner,however the exact mechanism remains obscure[39].
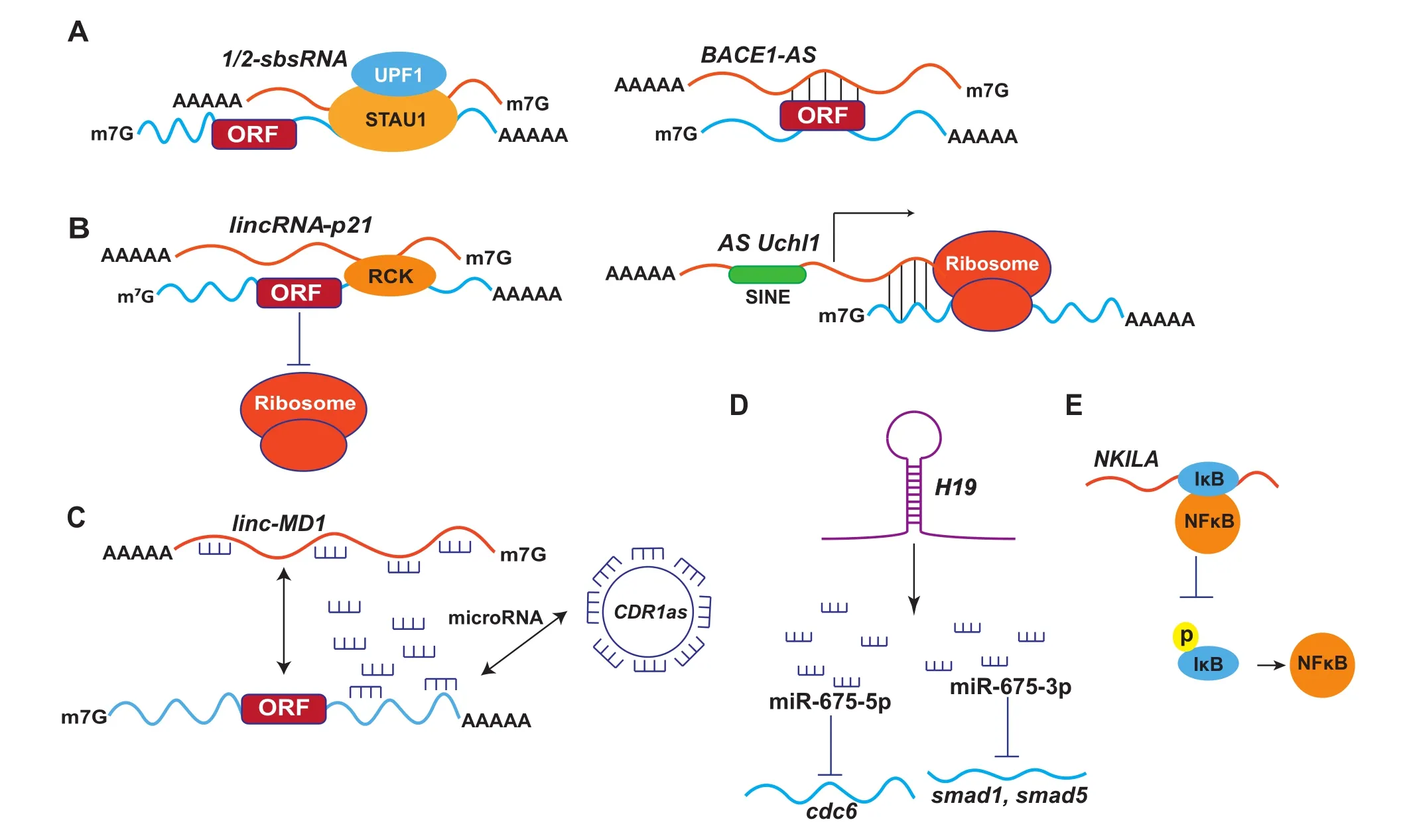
Figure 1 Known working models of cytoplasmic lncRNA function
Modulation of translation
Gene expression control at translational level plays a crucial role in multiple biological systems and provides valuable means for the spatiotemporal management of complex protein dynamics in eukaryotic cells[40-42]. Some lncRNAs also get involved in such regulation at the translational level,which can either repress(as exemplified for lincRNA-p21 below)or promote(as exemplified for AS Uchl1 below)translation.
lincRNA-p21
The human lincRNA-p21 is also known as tumor protein p53 pathway corepressor 1(Trp53cor1). lincRNA-p21 is~3.0 kb in length,and the encoding gene is located~15 kb upstream of p21/cdkn1a gene[23]. It is more abundant in cytoplasm compared to nucleus,known to co-distribute with ribosomes [43]. As a post-transcriptional modulator,lincRNA-p21 can negatively regulate the translation of CTNNB1(β-catenin)and JUNB transcripts by imperfectly base pairing at different sites in the coding and untranslated regions(both 5'and 3'UTRs)of CTNNB1(15 sites)and JUNB mRNAs(8 sites). When the level of Hu antigen R(HuR),a ubiquitous RNA binding protein,reduces,lincRNA-p21 becomes stable and interacts with its target transcripts including CTNNB1 and JUNB mRNAs. The resulting lincRNA-p21-mRNA complex can enhance the interaction between mRNAs and the translational repressors RCK as well as Fragile X mental retardation protein(FMRP). Consequently,translation of the target transcripts is repressed through reduced polysome sizes and ribosome drop-off(Figure 1B)[43,44].
AS Uchl1
A recent study reported the discovery of a spliced nuclearenriched antisense transcript(AS Uchl1)complementary to the mRNA that encodes mouse ubiquitin carboxy terminal hydrolase L1(Uchl1)[45]. Uchl1 is an enzyme specifically expressed in dopaminergic neurons[24,46,47]. The activity of the AS Uchl1 depends on the presence of a 73-nt overlapping sequence complementary with 5'end of Uchl1 mRNA andan embedded inverted SINEB2 repetitive element(Figure 1B)[45]. Under normal physiological conditions,AS Uchl1 is enriched in the nucleus,and upon rapamycin treatment,inhibition of mTORC1 triggers the transport of AS Uchl1 to the cytoplasm,which then targets the overlapping Uchl1 mRNA to active polysomes for cap-independent translation. Exact molecular mechanism as to how AS Uchl1 promotes the translation of Uchl1 mRNA under stress conditions is still elusive.
Competing endogenous RNAs
Coding and non-coding RNAs can regulate each other through their ability to compete for miRNA binding. lncRNAs harboring multiple binding sites of identical miRNA are called competing endogenous RNAs(ceRNAs)[48]. ceRNA can sequester miRNAs and therefore protect their target mRNAs from repression[49-53]. This activity was first discovered in Arabidopsis thaliana and later in mammals[53,54]. Multiple ceRNAs have been identified,and we present some as examples below.
HULC
Hepatocellular carcinoma(HCC)is one of the most fatal cancers[55]. Recent studies have indicated that a large number of lncRNAs are functionally deregulated in HCC[56-59]. Among these,highly up-regulated in liver cancer(HULC)is a novel mRNA-like ncRNA. It is present in the cytoplasm,spliced,polyadenylated,and resembles the mammalian LTR transposon 1A[60]. As reflected by its name,HULC is highly upregulated in HCC,and it is also detected in gastric cancer and colorectal carcinomas that metastasize to the liver[60-62]. The HULC gene resides on chromosome 6p24.3 in humans and is conserved in primates. It is about 1.6 kb in length and contains two exons. Expression of HULC gene in Hep3B cells can be up-regulated by the transcription factor cAMP responsive element binding protein(CREB). HULC acts as endogenous sponge of miR-372[63]. HULC binding to miR-372 reduces miRNA-mediated translational repression of protein kinase cAMP-activated catalytic subunit beta(PRKACB),one of the target genes of miR-372[63]. PRKACB can induce phosphorylation of CREB,which in turn stimulates HULC expression,thus forming a feedforward loop[63].
linc-MD1
linc-MD1 is a muscle-specific lncRNA,which is indispensable for the timing of muscle differentiation and plays an important role in myogenesis[64]. linc-MD1 acts as a natural decoy for two muscle-specific miRNAs,miR-133 and miR-135 (Figure 1C)[64]. Expression of mastermind-like-1(MAML1)is controlled by miR-133,and myocyte-specific enhancer factor 2C(MEF2C)is the target of miR-135[64]. MAML1 and MEF2C are important myogenic factors required for activation of muscle-specific genes. MEF2C binds to the promoter region of cardiac muscle genes and positively regulates the differentiation of muscle cells[65,66],while MAML1 acts as a transcription coactivator in some signal transduction pathways (such as Notch signaling)related to muscle differentiation[67]. With the depletion of linc-MD1,expression of both MAML1 and MEF2C is repressed,whereas over expression of linc-MD1 resulted in high levels of MAML1 and MEF2C. These observations argue for a direct competition between linc-MD1 and mRNAs for miRNA binding[64].
linc-RoR
The lncRNA regulator of reprogramming(linc-RoR)functions as microRNA(miRNA)sponge against miR-145. Interaction between linc-RoR and miR-145 prevents mRNA of some important transcription factors(TFs)like Oct4,sox2,and Nanog in human embryonic stem cells(hESCs)from miRNA-mediated regulation[68,69]. The expression of linc-RoR is positively correlated with the undifferentiated state of hESCs[69].
CDR1as and circSry
Recently,additional examples of ceRNA were found in circular RNAs(circRNAs),which represent a newly identified large class of lncRNAs[70-72]. circRNAs can be formed by backsplicing of the 5'end of an upstream exon with the 3'end of the same exon or a downstream exon. Although some circRNAs such as EIciRNAs are predominantly localized in the nucleus,circRNAs are generally cytoplasmic. circRNAs appear to be non-coding and lack the association with polysomes[71,73]. Two cytoplasmic circRNAs have been reported to act as miRNA sponge. The first one is the cerebellar degeneration-related protein 1 antisense transcript(CDR1as,also called ciRS-7),which is a sponge for miR-7(Figure 1C). CDR1as contains 74 miR-7 seed matches,out of which 63 are conserved in mammals[72]. The other one is a testisspecific circRNA encoded by the gene sex-determining region Y(circSry),which contains 16 putative binding sites for miR-138[71,72,74]. These two circRNAs may be special cases,and it may not be a general phenomenon for circRNAs to function as miRNA sponges[75].
Precursor of miRNAs
A genome-wide survey predicted that nearly 100 lncRNAs encode miRNAs[76]. These lncRNAs may not be predominantly cytoplasmic,but they may be processed in the nucleus and cytoplasm to give rise to functional miRNAs.
H19
H19 is one of the best known imprinting genes expressed from the maternal allele and required for proper muscle differentiation and muscle regeneration[77-79]. The H19 gene is present on chromosomes 11 and 7 in humans and mice,respectively [80,81]. There is no conserved ORF sequence in H19 RNA between mice and human. Although the H19 gene is imprinted paternally,the H19 RNA itself does not take part in imprinting mechanism[82]. Studies based on structure prediction suggest that H19 is a ncRNA,2.3-kb long,capped,spliced,and polyadenylated[82,83]. It is reported that H19 lncRNA acts as a molecular sponge for let-7 family of miRNAs in a HEK293 cell line[84]. Depleting H19 causes accelerated muscle differentiation,which can be recapitulated by let-7overexpression[84]. In the cytoplasm of undifferentiated multipotent mesenchymal C2C12 cells,H19 interacts with the K homology-type splicing regulatory protein(KSRP). Such binding favors KSRP-mediated destabilization of myogenin transcripts[85].
Besides the aforementioned roles of H19,exon 1 of H19 also gives rise to miR-675-3p and miR-675-5p(Figure 1D). miR-675-3p targets the gene encoding the anti-differentiation TFs smad1 and smad5,which are crucial components of the bone morphogenetic protein(BMP)pathway[86],whereas miR-675-5p targets the gene encoding DNA replication initiation factor Cdc6[86]. In this regard,H19 has a prodifferentiation function in primary myoblasts and regenerating skeletal muscles due to the resulting miR-675-3p and miR-675-5p[86,87]. H19 is also found to regulate placenta growth. Insulin like growth factor 2(Igf2),which is also targeted by miR-675-3p,is an important regulator of growth and is upregulated in H19-deficient placenta[88]. H19 is also found to modulate gastric cancer cell proliferation through miR-675,by targeting the gene encoding the tumor suppressor runt domain transcription factor1(RUNX1). Thus H19/miR-675 regulates the expression of RUNX1 to modulate gastric cancer[89].
linc-MD1(again)
We discussed linc-MD1 as a ceRNA before. However,linc-MD1 primary transcript also harbors the pri-miR-133b sequence. If cleaved by Drosha in the nucleus,linc-MD1 can give rise to a miRNA precursor. Recently,HuR protein is described as another component of linc-MD1 regulatory circuitry[90]. HuR is known to contribute to muscle differentiation[91]. HuR interacts with many coding and non-coding RNAs,indicating its pleiotropic RNA binding activity [92,93]. HuR binds to and favors linc-MD1 accumulation at the expense of miR-133 biogenesis. HuR also recruits miR-133 onto linc-MD1 in the cytoplasm,thereby reinforcing this regulatory circuitry. There is an inverse correlation between levels of HuR and miR-133b. HuR binds the base of the primiR-133b stem loop,and physically interferes with microprocessor activity[90]. Further investigations have to be carried out to answer how the processing and function of linc-MD1 are regulated either as the pri-miR-133b in the nucleus or as sponge for miR-133b when exported to the cytoplasm as an unprocessed transcript.
Regulation of protein modification
In the recent years,several lncRNAs are identified to modulate modifications of cytoplasmic proteins such as ubiquitination/ deubiquitination or phosphorylation/dephosphorylation.
lnc-DC
Expression of lnc-DC is almost exclusive to human conventional dendritic cells(DCs)[94]. lnc-DC could help activate STAT3 by binding to it in the cytoplasm,thus promoting the phosphorylation and preventing dephosphorylation of STAT3. Knockdown of lnc-DC inhibited the differentiation to the DC lineage as well as the functions of DCs[94].
NKILA
NF-κB interacting lncRNA(NKILA)binds directly to IκB and blocks IKK-induced IκB phosphorylation,thus inhibiting NF-κB activation(Figure 1E)[95]. The expression of NKILA is also upregulated by NF-κB. NKILA interacts with the NF-κB/IκB complex,and seems to keep the NF-κB pathway from over-activation and to suppress cancer metastasis[95].
Another role of lincRNA-p21
lincRNA-p21 was reported to regulate the ubiquitination of HIF-1α,a transcription factor crucial to hypoxia-induced effects such as Warburg effect[96]. lincRNA-p21 is induced by HIF-1α under hypoxia condition,and binds to both HIF-1α and von Hippel-Lindau tumor suppressor(VHL)protein. Such binding blocks the interaction between VHL and HIF-1α,thus inhibiting VHL-mediated ubiquitination of HIF-1α. This positive feedback loop between HIF-1α and lincRNA-p21 promotes glycolysis under hypoxia[96].
Perspectives
lncRNAs are recognized as major regulators in life events such as gene expression,cell differentiation,and tumorigenesis. In this article,we summarized the roles of some lncRNAs in the cytoplasm. lncRNAs can function in the posttranscriptional gene expression such as mRNA stability and translation. Through RNA-protein or RNA-RNA interaction,cytoplasmic lncRNAs could also serve as ceRNAs,miRNA precursors,or modulators of protein phosphorylation.
Recent findings have shown that certain transcripts previously-annotated as lncRNAs in fact can be translated to produce small bioactive peptides[97-100]. For instance,the conserved micropeptide myoregulin(MLN)was found to be encoded by a skeletal muscle-specific RNA,a previously putative lncRNA[100]. MLN shows structural and functional similarity with SERCA inhibitors,phospholamban and sarcolipin. Interacting directly with SERCA,MLN disrupts the Ca2+uptake into the sarcoplasmic reticulum[100]. Similarly,the endogenous 34-amino acid micropeptide dwarf open reading frame(DWORF)is encoded by another putative musclespecific lncRNA. DWORF enhances muscle performance by physically interacting with SERCA inhibitors such as phospholamban,sarcolipin,and MLN[100]. These examples demonstrated that some(although maybe limited in numbers)transcripts that are previously annotated as lncRNAs are actually coding,and thus should be considered as mRNAs. Given the vast amount of lncRNAs identified,and many of them are associated with noncoding functions,it is no doubt that more functions and functional working mechanisms are yet to be explored for the large number of cytoplasmic lncRNAs.
Regulation of lncRNA localization is important to coordinate their functions in the nucleus or in the cytoplasm. There should exist machinery either directly or indirectly to transport specific lncRNAs into the cytoplasm,and maybe further to special subcellular locations or complexes. The final localization,concentration,and functions of a specific lncRNA have to be fine tuned by the RNA biogenesis,transportation,degradation,and maybe even modifications. Substantial efforts are required to investigate these aspects.
A single lncRNA can have multiple roles. For example,both H19 and linc-MD1 can function as ceRNAs as well as precursors for miRNAs. How these different roles of the same lncRNA are coordinated remains to be addressed. On the other hand,there are undoubtedly more roles and functional mechanisms remain unknown for cytoplasmic lncRNAs. With the extensive investigations of the eukaryotic transcriptome by means of RNA sequencing,most of the lncRNAs including cytoplasmic ones may have already been described. Further studies on these lncRNAs may help to classify them into subclasses based on their biogenesis and functions.
Competing interests
The authors declare no competing interests.
Acknowledgments
This work is supported by the National Basic Research Program of China(973 Program;Grant No. 2015CB943000),the National Natural Science Foundation of China(Grant Nos. 91519333 and 31471225),and the Fundamental Research FundsfortheCentral Universities(GrantNo. WK2070000034).
References
[1]Mercer TR,Dinger ME,Mattick JS. Long noncoding RNAs: insights into functions. Nat Rev Genet 2009;10:155-9.
[2]Wilusz JE,Sunwoo H,Spector DL. Long noncoding RNAs: functional surprises fromthe RNA world. Genes Dev 2009;23:1494-504.
[3]Geisler S,Coller J. RNA in unexpected places: long non-coding RNA functions in diverse cellular contexts. Nat Rev Mol Cell Biol 2013;14:699-712.
[4]Hu S,Wu J,Chen L,Shan G. Signals from noncoding RNAs: unconventional roles for conventional pol III transcripts. Int J Biochem Cell Biol 2012;44:1847-51.
[5]Ramskold D,Wang ET,Burge CB,Sandberg R. An abundance of ubiquitously expressed genes revealed by tissue transcriptome sequence data. PLoS Comput Biol 2009;5:e1000598.
[6]Derrien T,Johnson R,Bussotti G,Tanzer A,Djebali S,Tilgner H,et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure,evolution,and expression. Genome Res 2012;22:1775-89.
[7]Moran VA,Perera RJ,Khalil AM. Emerging functional and mechanistic paradigms of mammalian long non-coding RNAs. Nucleic Acids Res 2012;40:6391-400.
[8]Batista PJ,Chang HY. Long noncoding RNAs: cellular address codes in development and disease. Cell 2013;152:1298-307.
[9]Mercer TR,Dinger ME,Sunkin SM,Mehler MF,Mattick JS. Specific expression of long noncoding RNAs in the mouse brain. Proc Natl Acad Sci U S A 2008;105:716-21.
[10]Wilusz JE. Long noncoding RNAs: Re-writing dogmas of RNA processing and stability. Biochim Biophys Acta 2016;1859: 128-38.
[11]Zhao J,Sun BK,Erwin JA,Song JJ,Lee JT. Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science 2008;322:750-6.
[12]Hung T,Wang Y,Lin MF,Koegel AK,Kotake Y,et al. Extensive and coordinated transcription of noncoding RNAs within cell-cycle promoters. Nat Genet 2011;43:621-9.
[13]Tripathi V,Ellis JD,Shen Z,Song DY,Pan Q,Watt AT,et al. The nuclear-retained non-coding RNA MALAT1 regulates alternative splicing by modulating SR splicing factor phosphorylation. Mol Cell 2010;39:925-38.
[14]Bernard D,Prasanth KV,Tripathi V,Colasse S,Nakamura T,Xuan Z,et al. A long nuclear-retained non-coding RNA regulates synaptogenesis by modulating gene expression. EMBO J 2010;29:3082-93.
[15]Ingolia NT,Lareau LF,Weissman JS. Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell 2011;147:789-802.
[16]Montes M,Nielsen MM,Maglieri G,Jacobsen A,Hojfeldt J,Agrawal-Singh S,et al. The lncRNA MIR31HG regulates p16INK4A expression to modulate senescence. Nat Commun 2015;6:6967.
[17]Gong C,Maquat LE. LncRNAs transactivate STAU1- mediated mRNA decay by duplexing with 39 UTRs via Alu elements. Nature 2011;470:284-8.
[18]Kim YK,Furic L,Parisien M,Major F,DesGroseillers L,Maquat LE,et al. Staufen1 regulates diverse classes of mammalian transcripts. EMBO J 2007;26:2670-81.
[19]Hollander MC,Alamo I,Fornace Jr AJ. A novel DNA damage inducible transcript,gadd7,inhibits cell growth,but lacks a protein product. Nucleic Acids Res 1996;24:1589-93.
[20]Fornace Jr AJ,Alamo Jr I,Hollander MC. DNA damage inducible transcripts in mammalian cells. Proc Natl Acad Sci U S A 1988;85:8800-4.
[21]Brookheart RT,Michel CI,Listenberger LL,Ory DS,Schaffer JE. The non-coding RNA gadd7 is a regulator of lipid-induced oxidative and endoplasmic reticulum stress. J Biol Chem 2009;284:7446-54.
[22]Rinn JL,Kertesz M,Wang JK,Squazzo SL,Xu X,Brugmann SA,et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 2007;129:1311-23.
[23]Huarte M,Guttman M,Feldser D,Garber M,Koziol MJ,Kenzelmann-Broz D,et al. A large intergenic noncoding RNA induced by p53 mediates global gene repression in the p53 response. Cell 2010;142:409-19.
[24]Liu H,Wei L,Tao Q,Deng H,Ming M,Xu P,et al. Decreased NURR1 and PITX3 gene expression in Chinese patients with Parkinson’s disease. Eur J Neurol 2012;19:870-5.
[25]Buratti E,Baralle FE. Multiple roles of TDP-43 in gene expression,splicing regulation,and human disease. Front Biosci 2008;13:867-78.
[26]Cohen TJ,Lee VM,Trojanowski JQ. TDP-43 functions and pathogenic mechanisms implicated in TDP-43 proteinopathies. Trends Mol Med 2011;17:659-67.
[27]Lee MH,Yang HY. Regulators of G1 cyclin-dependent kinases and cancers. Cancer Metastasis Rev 2003;22:435-49.
[28]Ayala YM,Misteli T,Baralle FE. TDP-43 regulates retinoblastoma protein phosphorylation through the repression of cyclin dependent kinase 6 expression. Proc Natl Acad Sci U S A 2008;105:3785-9.
[29]Pang KC,Frith MC,Mattick JS. Rapid evolution of noncoding RNAs: lack of conservation does not mean lack of function. Trends Genet 2006;22:1-5.
[30]Hellwig S,Bass BL. A starvation-induced noncoding RNA modulates expression of Dicer-regulated genes. Proc Natl Acad Sci U S A 2008;105:12897-902.
[31]Kabashi E,Valdmanis PN,Dion P,Spiegelman D,McConkey BJ,Velde CV. TARDBP mutations in individuals with sporadic andfamilial amyotrophic lateral sclerosis. Nat Genet 2008;40:572-4.
[32]Sreedharan J,Blair IP,Tripathi VB,Hu X,Vance C,Rogelj B,et al. TDP-43 mutations in familial and sporadic amyotrophic lateral sclerosis. Science 2008;319:1668-72.
[33]Faghihi MA,Modarresi F,Khalil AM,Wood DE,Sahagan BG,Morgan TE. Expression of a noncoding RNA is elevated in Alzheimer’s disease and drives rapid feed-forward regulation of beta-secretase. Nat Med 2008;14:723-30.
[34]Faghihi MA,Ming Z,Jia H,Farzaneh M,Van der Brug MP,Michael AN,et al. Evidence for natural antisense transcriptmediated inhibition of microRNA function. Genome Biol 2010;11:R56.
[35]Kretz M,Siprashvili Z,Chu C,Webster DE,Zehnder A,Qu K,et al. Control of somatic tissue differentiation by the long noncoding RNA TINCR. Nature 2013;493:231-5.
[36]Wan D,Gong Y,Qin W,Zhang P,Li J,Wei L,et al. Large-scale cDNA transfection screening for genes related to cancer development and progression. Proc Natl Acad Sci US A 2004;101:15724-9.
[37]Kiebler MA,Hemraj I,Verkadi P,Kohrmann M,Fortes P,Marion RM,et al. The mammalian staufen protein localizes to the somatodendritic domain of cultured hippocampal neurons: implications for its involvement in mRNA transport. J Neurosci 1999;19:288-97.
[38]Dugre´-Brisson S,Elvira G,Boulay K,Chatel-Chaix L,Mouland AJ,DesGroseillers L. Interaction of Staufen1 with the 5'end of mRNA facilitates translation of these RNAs. Nucleic Acids Res 2005;33:4797-812.
[39]Kretz M. TINCR,staufen1,and cellular differentiation. RNA Biol 2013;10:1597-601.
[40]Gray NK,Wickens M. Control of translation initiation in animals. Annu Rev Cell Dev Biol 1998;14:399-458.
[41]Sonenberg N,Hershey JWB,Mathews MB. Translational control of gene expression. Cold Spring Harbor,New York: Cold Spring Harbor Laboratory Press;2000. p. 1020.
[42]Gebauer F,Hentze MW. Molecular mechanisms of translational control. Nat Rev Mol Cell Biol 2004;5:827-35.
[43]Yoon JH,Abdelmohsen K,Srikantan S,Yang X,Martindale JL,De S,et al. LincRNA-p21 suppresses target mRNA translation. Mol Cell 2012;47:648-55.
[44]Chu CY,Rana TM. Translation repression in human cells by microRNA-induced gene silencing requires RCK/p54. PLoS Biol 2006;4:e210.
[45]Carrieri C,Cimatti L,Biagioli M,Beugnet A,Zucchelli S,Fedele S,et al. Long non-coding antisense RNA controls Uchl1 translation through an embedded SINEB2 repeat. Nature 2012;491:454-7.
[46]Liu Y,Fallon L,Lashuel HA,Liu Z,Lansbury Jr PT. The UCHL1 gene encodes two opposing enzymatic activities that affect alpha-synuclein degradation and Parkinson’s disease susceptibility. Cell 2002;111:209-18.
[47]Setsuie R,Wada K. The functions of UCH-L1 and its relation to neurodegenerative diseases. Neurochem Int 2007;51:105-11.
[48]Salmena L,Poliseno L,Tay Y,Pandolfi PP. The ceRNA hypothesis: the Rosetta stone of a hidden RNA language. Cell 2011;146:353-8.
[49]Karreth FA,Tay Y,Perna D,Ala U,Tan SM,Rust AG,et al. In vivo identification of tumor-suppressive PTEN ceRNAs in an oncogenic BRAF-induced mouse model of melanoma. Cell 2011;147:382-95.
[50]Tay Y,Kats L,Salmena L,Weiss D,Tan SM,Ala U,et al. Coding-independent regulation of the tumor suppressor PTEN by competing endogenous mRNAs. Cell 2011;147:344-57.
[51]Peng W,Si S,Zhang Q,Li C,Zhao F,Wang F,et al. Long noncoding RNA MEG3 functions as a competing endogenous RNA to regulate gastric cancer progression. J Exp Clin Cancer Res 2015;34:79-89.
[52]Chen MT,Lin HS,Shen C,Ma YN,Wang F,Zhao HL,et al. PU.1-regulated long noncoding RNA lnc-MC controls human monocyte/macrophage differentiation through interaction with microRNA 199a-5p. Mol Cell Biol 2015;35:3212-24.
[53]Franco-Zorrilla JM,Valli A,Todesco M,Mateos I,Puga MI,Rubio-Somoza I,et al. Target mimicry provides a new mechanismfor regulation of microRNAactivity. Nat Genet 2007;39:1033-7.
[54]Poliseno L,Salmena L,Zhang J,Carver J,Haveman J,Pandolfi PP,et al. A coding-independent function of gene and pseudogene mRNAs regulates tumor biology. Nature 2010;465:1033-8.
[55]Bosch FX,Ribes J,Cle´ries R,Diaz M. Epidemiology of hepatocellular carcinoma. Clin Liver Dis 2005;9:191-211.
[56]Matouk IJ,DeGroot N,Mezan S,Ayesh S,Abu-lail R,Hochberg A,et al. The H19 non-coding RNA is essential for human tumor growth. PLoS One 2007;9:e845.
[57]Tsang WP,Kwok TT. Riboregulator H19 induction of MDR1-associated drug resistance in human hepatocellular carcinoma cells. Oncogene 2007;26:4877-81.
[58]Ventura A,Jacks T. MicroRNAs and cancer: short RNAs go a long way. Cell 2009;136:586-91.
[59]Qin R,Chen Z,Ding Y,Hao J,Hu J,Guo F. Long non-coding RNA MEG3 inhibits the proliferation of cervical carcinoma cells through the induction of cell cycle arrest and apoptosis. Neoplasma 2013;60:486-92.
[60]Panzitt K,Tschernatsch MM,Guelly C,Moustafa T,Stradner M,Strohmaier HM,et al. Characterization of HULC,a novel gene with striking up-regulation in hepatocellular carcinoma,as noncoding RNA. Gastroenterology 2007;132:330-42.
[61]Matouk IJ,Abbasi I,Hochberg A,Galun E,Dweik H,Akkawi M. Highly upregulated in liver cancer noncoding RNA is overexpressed in hepatic colorectal metastasis. Eur J Gastroenterol Hepatol 2009;21:688-92.
[62]Zhao Y,Guo Q,Chen J,Hu J,Wang S,Sun Y. Role of long non-coding RNA HULC in cell proliferation,apoptosis and tumor metastasis of gastric cancer: a clinical and in vitro investigation. Oncol Rep 2014;31:358-64.
[63]Wang J,Liu X,Wu H,Ni P,Gu Z,Qiao Y,et al. CREB upregulates long non-coding RNA,HULC expression through interaction with microRNA-372 in liver cancer. Nucleic Acids Res 2010;38:5366-83.
[64]Cesana M,Cacchiarelli D,Legnini I,Santini T,Sthandier O,Chinappi M,et al. A long noncoding RNA controls muscle differentiation by functioning as a competing endogenous RNA. Cell 2011;147:358-69.
[65]Lilly B,Zhao B,Ranganayakulu G,Paterson BM,Schulz RA,Olson EN. Requirement of MADS domain transcription factor D-MEF2 for muscle formation in Drosophila. Science 1995;267:688-93.
[66]Lin Q,Schwarz J,Bucana C,Olson EN. Control of mouse cardiac morphogenesis and myogenesis by transcription factor MEF2C. Science 1997;276:1404-7.
[67]Shen H,McElhinny AS,Cao Y,Gao P,Liu J,Bronson R,et al. The Notch coactivator,MAML1,functions as a novel coactivator for MEF2C-mediated transcription and is required for normal myogenesis. Genes Dev 2006;20:675-88.
[68]Loewer S,Cabili MN,Guttman M,Loh YH,Thomas K,Park IH. Large intergenic non-coding RNA-RoR modulates reprogramming of human induced pluripotent stem cells. Nat Genet 2010;42:1113-7.
[69]Wang Y,Xu Z,Jiang J,Xu C,Kang J,Xiao L,et al. Endogenous miRNA sponge lincRNA-RoR regulates Oct4,Nanog,and Sox2 in human embryonic stem cell self-renewal. Dev Cell 2013;25:69-80.
[70]Salzman J,Gawad C,Wang PL,Lacayo N,Brown PO. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS One 2012;7:e30733.
[71]Hansen TB,Jensen TI,Clausen BH,Bramsen JB,Finsen B,Damgaard KC,et al. Natural RNA circles function as efficient microRNA sponges. Nature 2013;495:384-8.
[72]Memczak S,Jens M,Elefsinioti A,Torti F,Krueger J,Rybak A. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 2013;495:333-8.
[73]Li Z,Huang C,Bao C,Chen L,Lin M,Wang X,et al. Exonintron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol 2015;22:256-64.
[74]Capel B,Swain A,Nicolis S,Hacker A,Walter M,Koopman P,et al. Circular transcripts of the testis-determining gene Sry in adult mouse testis. Cell 1993;73:1019-30.
[75]Guo JU,Agarwal V,Guo H,Bartel DP. Expanded identification and characterization of mammalian circular RNAs. Genome Biol 2014;15:409.
[76]He S,Su H,Liu C,Skogerbo G,He H,He D,et al. MicroRNA-encoding long non-coding RNAs. BMC Genomics 2008;9:236.
[77]Pachnis V,Belayew A,Tilghman SM. Locus unlinked to afetoprotein under the control of the murine raf and Rif genes. Proc Natl Acad Sci U S A 1984;81:5523-7.
[78]Poirier F,Chan CT,Timmons PM,Robertson EJ,Evans MJ,Rigby PW. The murine H19 gene is activated during embryonic stem cell differentiation in vitro and at the time of implantation in the developing embryo. Development 1991;113:1105-14.
[79]Davis RL,Weintraub H,Lassar AB. Expression of a single transfected cDNA converts fibroblasts to myoblasts. Cell 1987;51:987-1000.
[80]Bartolomei MS,Zemel S,Tilghman SM. Parental imprinting of the mouse H19 gene. Nature 1991;351:153-5.
[81]Zhang Y,Tycko B. Monoallelic expression of the human H19 gene. Nat Genet 1992;1:40-4.
[82]Brannan CI,Dees EC,Ingram RS,Tilghman SM. The product of the H19 gene may function as an RNA. Mol Cell Biol 1990;10:28-36.
[83]Juan V,Crain C,Wilson C. Evidence for evolutionarily conserved secondary structure in the H19 tumor suppressor RNA. Nucleic Acids Res 2000;28:1221-7.
[84]Kallen AN,Zhou XB,Xu J,Qiao C,Ma J,Yan L,et al. The imprinted H19 lncRNA antagonizes let-7 microRNAs. Mol Cell 2013;52:101-12.
[85]Giovarellia M,Bucci G,Ramos A,Bordo D,Wilusz CJ,Chen CY,et al. H19 long noncoding RNA controls the mRNA decay promoting function of KSRP. Proc Natl Acad Sci U S A 2014;111:E5023-8.
[86]Dey BK,Karl P,Anindya D. The H19 long noncoding RNA gives rise to microRNAs miR-675-3p and miR-675-5p to promote skeletal muscle differentiation and regeneration. Genes Dev 2014;28:491-501.
[87]Cai X,Cullen BR. The imprinted H19 noncoding RNA is a primary microRNA precursor. RNA 2007;13:313-6.
[88]Keniry A,Oxley D,Monnier P,Kyba M,Dandolo L,Smits G,et al. The H19 lincRNA is a developmental reservoir of miR-675 that suppresses growth and Igf1r. Nat Cell Biol 2012;14:659-65.
[89]Zhuang M,Gao W,Xu J,Wang P,Shu Y. The long non-coding RNA H19 derived miR-675 modulates human gastric cancer cell proliferation by targeting tumor suppressor RUNX1. Biochem Biophys Res Commun 2014;448:315-22.
[90]Legnini I,Morlando M,Mangiavacchi A,Fatica A,Bozzoni I. A feedforward regulatory loop between HuR and the long noncoding RNA linc-MD1 controls early phases of myogenesis. Mol Cell 2014;53:506-14.
[91]Von Roretz C,Beauchamp P,Di Marco S,Gallouzi IE. HuR and myogenesis: being in the right place at the right time. Biochim Biophys Acta 2011;1813:1663-7.
[92]Lebedeva S,Jens M,Theil K,Schwanha¨usser B,Selbach M,Landthaler M,et al. Transcriptome-wide analysis of regulatory interactions of the RNA-binding protein HuR. Mol Cell 2011;43:340-52.
[93]Mukherjee N,Corcoran DL,Nusbaum JD,Reid DW,Georgiev S,Hafner M,et al. Integrative regulatory mapping indicates that the RNA-binding protein HuR couples pre-mRNA processing and mRNA stability. Mol Cell 2011;43:327-39.
[94]Wang P,Xue Y,Han Y,Lin L,Wu C,Xu S,et al. The STAT3-binding long noncoding RNA lnc-DC controls human dendritic cell differentiation. Science 2014;344:310-3.
[95]Liu B,Sun L,Liu Q,Gong C,Yao Y,Lv X,et al. A cytoplasmic NF-κB interacting long noncoding RNA blocks IκB phosphorylation and suppresses breast cancer metastasis. Cancer Cell 2015;27:370-81.
[96]Yang F,Zhang H,Mei Y,Wu M. Reciprocal regulation of HIF-1α and lincRNA-p21 modulates the Warburg effect. Mol Cell 2014;53:88-100.
[97]Kondo T,Plaza S,Zanet J,Benrabah E,Valenti P,Hashimoto Y,et al. Small peptides switch the transcriptional activity of ShavenbabyduringDrosophilaembryogenesis. Science 2010;329:336-9.
[98]Andrews SJ,Rothnagel JA. Emerging evidence for functional peptides encoded by short open reading frames. Nat Rev Genet 2014;15:193-204.
[99]Anderson DM,Anderson KM,Chang CL,Makarewich CA,Nelson BR,McAnally JR,et al. A micropeptide encoded by a putative long noncoding RNA regulates muscle performance. Cell 2015;160:595-606.
[100]Nelson BR,Makarewich CA,Anderson DM,Winders BR,Troupes CD,Wu F,et al. A peptide encoded by a transcript annotated as long noncoding RNA enhances SERCA activity in muscle. Science 2016;351:271-5.
ORIGINAL RESEARCH
*Corresponding author.
杂志排行
Genomics,Proteomics & Bioinformatics的其它文章
- Genomic,Lipidomic,and Metabolomic Analysis of Cyclooxygenase-null Cells:Eicosanoid Storm,Cross Talk,and Compensation by COX-1
- LVTree Viewer: An Interactive Display for the All-Species Living Tree Incorporating Automatic Comparison with Prokaryotic Systematics
- Similarity Estimation Between DNA Sequences Based on Local Pattern Histograms of Binary Images
- Up-regulation of Long Non-coding RNA TUG1 in Hibernating Thirteen-lined Ground Squirrels
- Call for Papers Special Issue on“Genome Stability’’
- Call for Papers Special Issue on“Computational Cardiology’’
