A quadratic linear-parabolic model-based EEG classification to detect epileptic seizures
2020-09-21AntonioQuinteroRincCarlosGianoHadjBatatia
Antonio Quintero-Rincón, Carlos D'Giano, Hadj Batatia
1Epilepsy and Telemetry Integral Center, Foundation for the Fight against Pediatric Neurological Disease, Montañeses 2325, Buenos Aires C1428AQK, Argentina;2Computer Science Research Institute of Toulouse–National Polytechnic Institute of Toulouse, University of Toulouse,Toulouse, Cedex 7 B.P. 7122-31071, France.
Abstract The two-point central difference is a common algorithm in biological signal processing and is particularly useful in analyzing physiological signals. In this paper, we develop a model-based classification method to detect epileptic seizures that relies on this algorithm to filter electroencephalogram (EEG) signals. The underlying idea was to design an EEG filter that enhances the waveform of epileptic signals. The filtered signal was fitted to a quadratic linear-parabolic model using the curve fitting technique. The model fitting was assessed using four statistical parameters, which were used as classification features with a random forest algorithm to discriminate seizure and non-seizure events. The proposed method was applied to 66 epochs from the Children Hospital Boston database. Results showed that the method achieved fast and accurate detection of epileptic seizures, with a 92% sensitivity, 96% specificity, and 94.1% accuracy.
Keywords: two-point central difference, curve fitting, parabolic curves, epilepsy, electroencephalogram, random forest
Introduction
Epilepsy is a neurological disease that affects people of all ages. It is characterized by unpredictable seizures resulting from the hyperexcitability of neurons. The electroencephalogram (EEG) is the predominant modality to study cerebral activity. The analysis of EEG signals is nearly completely dependent on visual inspection by the physician to quantify or qualify the morphology of waves. Their goal is the identification and classification of abnormal patterns in order to provide aid for an epilepsy diagnosis.
This study proposes a simple, fast, and adaptable method, implementable in real-time, to help physicians visually inspect EEG signals for epileptic seizure detection. The proposed method fits within the framework of model-based classification. It is based on the statistical parameters obtained from fitting a quadratic parabolic model to specifically filtered and transformed EEG signals. Precisely, the two-point central difference algorithm is used to build a filter for the EEG signals. The filtered signal is subsequently represented using a quadratic linear-parabolic model,using the curve fitting approach. Four statistical parameters are then calculated to characterize the model fitting. These parameters are considered as a feature-vector in the classification stage.
The literature abounds with research work dealing with the detection of epileptic seizure onset. Various methods and techniques have been proposed for this purpose. Many existing methods fit within the large framework of the feature-based machine learning approach.
The two-point central difference algorithm has proven useful in analyzing physiological signals, with various medical applications reported in the literature.A typical work has been reported by Freiet alto detect epileptiform discharges in muscles in EEG signals[1]. This algorithm is usually used to estimate the derivative of a function, giving an estimate valid only over a limited frequency range. Here, our idea was to use this algorithm to design a filter that enhances the frequency range of EEG signals corresponding to brain activity characteristic of epileptic seizures. To single out the related waveform,we transform the filtered signal into a quadratic form.A quadratic linear-parabolic model is then fitted to the transformed signal, using the curve fitting technique[2–3].The fitting is assessed using four statistical parameters(weighted sum of squared residuals, R-square,adjusted R-square, and root mean squared error). In order to show that our quadratic linear-parabolic model is pertinent to characterize epileptic seizures,we develop a classification method where the modelfitting parameters are used as features. We show that these features are good biomarkers of epileptic seizures.
The curve fitting technique adopted here is widely used in signal processing in general, and biomedical applications in particular. It has also been applied a few times in the domain of EEG and epilepsy. Orhanet alused the polynomial curve fitting method to fit a probability density function (PDF) of EEG signals discretized using equal frequency binning. The method has been applied to epileptic seizure detection using the comparison of PDFs of seizures and nonseizures. Second-order Fourier curve fitting has been used to smooth EEG signals in a pre-processing stage of a model-based method to predict epileptic seizures[4]. A spline curve fitting to EEG data was proposed to detect phase cone patterns to characterize epileptic activity[5]. Linear curve fitting was applied to interictal heartbeat intervals in order to combine EEG and ECG signals with the purpose of characterizing modulation patterns in patients with drug-resistant epilepsy[6]. More recently, a first-order exponential function has been fit to voltage-gated sodium channel type 2 to model the conductance-voltage curves in neonatal epileptic subjects[7].
Supervised machine learning methods have been widely used to detect seizures in EEG, with support vector machine (SVM) and K-Nearest Neighbor(KNN) being the most popular. SVM classifiers have been used with various features extracted from spectral and entropy analysis[8], matching pursuit algorithm[9], tensor discriminant analysis[10],multifractal detrended fluctuation analysis[11], and cross-bispectrum analysis[12]. While KNN classifiers have been developed, among many others, with features from the fractal dimension[13], and non-linear dimension reduction of frequency domain parameters[14]. In addition to these two popular approaches, Acharyaet alhave recently been the first to develop a seizure detection method using a convolutional neural network[15].
In general, these methods have good performance thanks to the advanced signal processing techniques they rely on. However, they consequently have high computational costs[16]. In this work, we combine for the first time the two-point difference algorithm and the quadratic linear-parabolic model to design an original model-based statistical classification method to detect epileptic seizures in EEG. The proposed feature-vector is simple and fast to calculate from the quadratic model. Our classification method is based on bootstrap-aggregated (bagged) decision trees. This approach combines results from several decision trees to overcome the overfitting effect due to the variability in the training data[17]. Using the different subset of features (subspace sampling) to build the weak-tree classifiers reduces the training time[18], and there are some recent works in seizure classification using EEG[19–22].
Materials and methods
Data
For the experimentation purpose, we considered a dataset from the Children's Hospital Boston database[23–24]which consists of 22 EEG recordings from pediatric subjects with intractable seizures. All signals were sampled at 256 Hz with a 16-bit resolution by using the International 10-20 system.The set of recordings lasted on average 35 minutes for 30 subjects in total, 2 hours for 4 subjects, and 12 hours for 2 other subjects. Taken together the recordings account for 60 hours of EEG recordings and 139 seizures. No distinctions regarding the types of seizure onsets were considered. The data contains focal, lateral, and generalized seizure onsets.Furthermore, the recordings were made in a routine clinical environment, so non-seizure activity and artifacts such as head/body movement, chewing,blinking, early stages of sleep, and electrode pops/movements were present in the data. In this work, we have used 66 epochs from 9 different subjects (Table 1). Each recording contains a seizure event, whose onset time has been labeled by an expert neurologist. The duration has an average of 79.4 seconds. Moreover, for each seizure segment, the neurologist also selected one adjacent non-seizure signal segment before and after the seizure, of the same length to represent healthy brain activity. In total, we considered 33 seizures and 33 non-seizure events. Non-seizure control events have been chosen just before seizures. This choice is justified for two reasons. First, we are interested in identifying seizure onset, therefore it is the most important to distinguish its signal from preceding normal activity. Second, the EEG activity following a seizure remains chaotic for a long time, and comparing it to seizure onset is not pertinent. The selected signals had the same montage by using 23 scalp EEG channels.
Methodology
SeaX∈RNxMdenotes the matrix ofMEEG signalsxm∈RNx1, measured simultaneously on different channels and atNdiscrete-time instants. The proposed methodology is composed of five stages. The first stage divides the original signalXinto a set of nonoverlapping 1-second segments using a rectangular sliding window Ω =Ω0(ω−(W−1)/2) with 0≤ω≤W–1, so thatX[n]=Ω[n]X. In the second stage, the two-point central difference algorithm is used to estimate the coefficientsb[k]of the filter for each 1-second segmentX[n]. The signal is then convolved by the resulting filterX˜ [n]=X[n]×b[k]. The third stage fits a quadratic linear-parabolic model to the filtered signal and estimates the associated parameters. The fourth stage calculates four statistical parameters for each EEG segment, namely the weighted sum of squared residuals (ζ) the R-square value (ϕ), the adjusted R-square value (σ) and the root means squared error (ψ), to assess the quadratic model fitting. Finally, in stage five, the feature-vectorρ=[ζ,ϕ,σ,ψ]associated with each segment is classified using a random forest classifier to discriminate between seizure and non-seizure.
Two-point central difference algorithm
The principle of the two-point central difference algorithm consists in subtracting non-adjacent,regularly spaced, and pairs of points. The objective is to extract a slope fromX[n]:
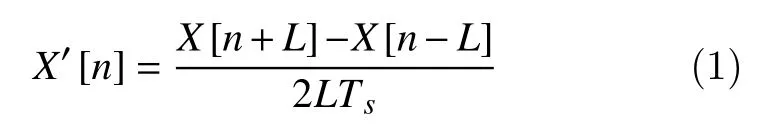
whereLis called the skip factor that defines the distance between points, andTSis the sample interval scaling. Taking the Z-transform of both sides, we obtain
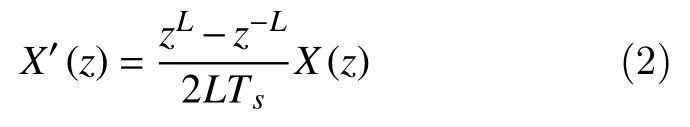
The transfer function of (2) is given by
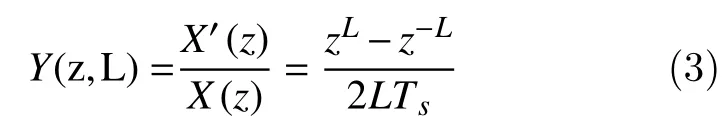
Thus, the frequency response is obtained by replacingzbyejΩ
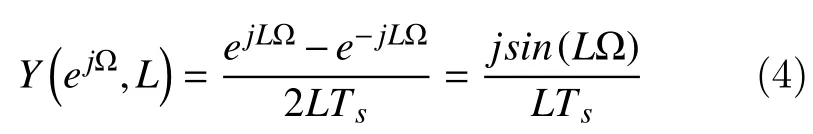
Using a symmetric FIR (finite impulse response)filter, the two-point central difference algorithm is based on an impulse function containing two coefficients of equal but opposite sign spacedLpoints apart with the following coefficients
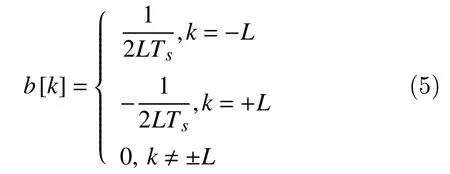
Note that, FIR filters are free from stability problems and they cause no time delay and no phase distortion within the pass-band. A convolution operation is estimated between each 1-second segmentX[n]and the coefficientsb[k]to obtain a new filtered signal in a bandwidth frequency until 50 Hz:
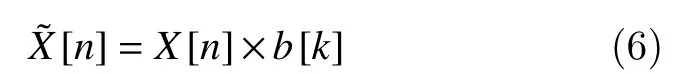
Note that, this is the effective bandwidth of the filter where the EEG activity has an important clinical relevance[25]. This allows us to automatically reject the high line-noise artifacts greater than 50 Hz asL=Fs/5,as the sampling rateFsis the 256 Hz. Note thatLis rounded. A comprehensive treatment of the mathematical properties of the two-point central difference algorithm is referenced[26–28].
Quadratic linear-parabolic model
We propose to fit a quadratic linear-parabolic model to the filtered signal. Precisely, the model is fitted to the couplewhereis the complete filtered signal, obtained by concatenating the segments given in equation (6).
The idea is to fit the signal with a curve of the form:
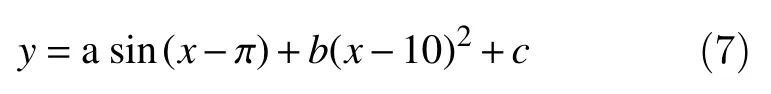
The fitting is performed using the least-squares method to estimate the three parameters a, b, c[29–31].
Model-fitting statistical parameters
To assess the fitting of the curve given in equation(7), the following four statistical parameters are estimated. Note thatyis the observed data,is the predicted value using the quadratic linear-parabolic model, andis the mean of the observed data.
Weighted sum of squared residuals (ζ): It is used to measure the total deviation of the response values from the predicted values, and is defined as

The weights allow taking into consideration the different uncertainties of the measurements and are calculated as follows

whereσ2is the variance.
R-square value (ϕ): It is the square of the correlation between the response values and the predicted response values, and is defined as
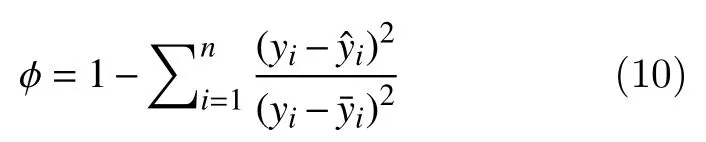
Adjusted R-square value (σ): It adjusts the R-square residual degrees of freedom, and is defined as
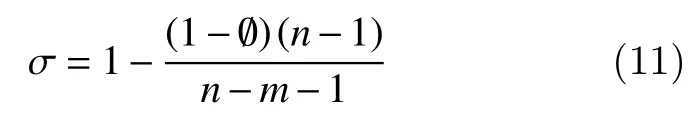
wherenis the number of response values andnis the estimated fitted coefficients from the response values. A value ofσcloser to 1 indicates a better fit.
Root mean squared error (ψ): It is an estimate of the standard deviation of the random component in the data and is defined as

Random forest classification
In this section, we present a classification method to identify seizures. The four statistical parameters presented above are used as classification features. A random forest classification technique is adopted.Random forest is an ensemble learning technique that combines the Bagging algorithm and the random subspace method using decision trees as the base classifier. The goal is to discriminate between seizure and non-seizure events, specifically the seizure onset through the feature predictor vectorρ=[ζ,ϕ,σ,ψ]associated with each EEG segment. The idea is to design a prediction functionf(ρ) to estimate the binary response for two classes, Γ={γ1,γ0}, whereγ1=1 for a seizure event andγ0=0 for a non-seizure event. A loss functionL(Γ,f(ρ)) determines the prediction function as follows:
whereEρ,Γdenotes the expectation with respect to an unknown joint distributionofρand Γ.For the classification, the next zero-one loss function is used
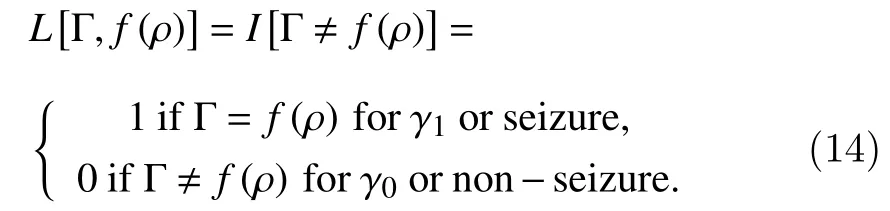
The ensemble learning approach constructsfby usingJdecision-tree base-classifiers
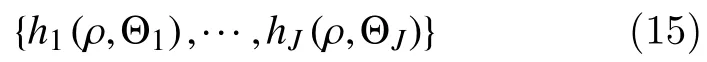
where Θj,j=1,···,J, is an independent collection of random variables. These base classifiers are combined by majority vote, so that
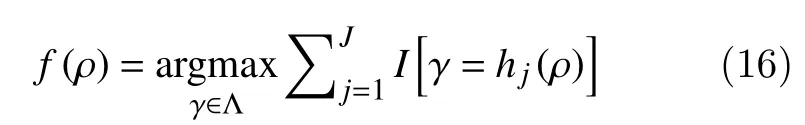
Consider a dataset D={d1,d2,···,dN,} withdi=(ρi,ci), whereciis a class label,ρdenotes the vector predictor andγithe response. For a particular realizationθjof Θj, the fitted tree is denotedThe random forest classifier uses random subspace in two ways. First, each tree is fitted to an independent bootstrap sample from the original data.The randomization involved in bootstrap sampling gives one part of Θj. Second, the best split is retained for allρpredictors according to the randomly selected subset ofmpredictor variables.
The randomization used to sample the predictors gives the remaining part of Θj. The resulting class predicted is by the majority combination of unweighted voting of the trees (Algorithm 1). A comprehensive treatment of the properties of random forest classifier are referenced[17–18,32].
Results
In this section, we evaluated the proposed methodology using the Children Hospital Boston database, which presented in the Material and methods section.
Model fitting
This section presents the results of the quadratic linear-parabolic model fitting.Table 2presents the three coefficients (a,b,c) estimated according to equation (7). Coefficients for seizures and nonseizures are shown separately, for illustration purposes only. This allows us to note the significant difference between values corresponding to the two classes. The actual quadratic linear-parabolic model is fitted to the entire signal, without distinction of seizure and nonseizure.

?
Fig. 1shows the quadratic linear-parabolic curves obtained for seizure and non-seizure segments,respectively. One notices that the proposed model (7)provides a visually precise fit to the EEG waveform.In addition, the prediction bounds have a small uncertainty throughout the entire data range, hence new observations should be predicted with high accuracy.
Classification
The dataset described above has been used to train the classifier according to the proposed method. The capacity of the proposed classification scheme to discriminate between seizure and non-seizure events in order to detect the seizure onset in EEG signals has been assessed. InFig. 2, we can observe the great difficulty in discriminating between seizure and nonseizure in EEG raw data. The start and end of the seizure in this EEG signal were labeled by the neurologist using two lines. The first line divides the EEG signal at 81 second (onset) and the second at 162 second (offset).
Table 3reports the four statistical parameters,calculated for all seizure and non-seizure segments.The weighted sum of squared residuals (ζ) and the root mean squared error (ψ) show much larger values for seizure events with respect to non-seizure events,which suggests that these features can be used to discriminate between seizure and non-seizures in EEG signals. While R-square (ϕ) and adjusted R-square (σ)have values close to one for both types of events,which suggests that the model has high accuracy in the fit. These four parameters show good potential todiscriminate between seizure and non-seizures in EEG signals. This can be visually corroborated inFig. 1.
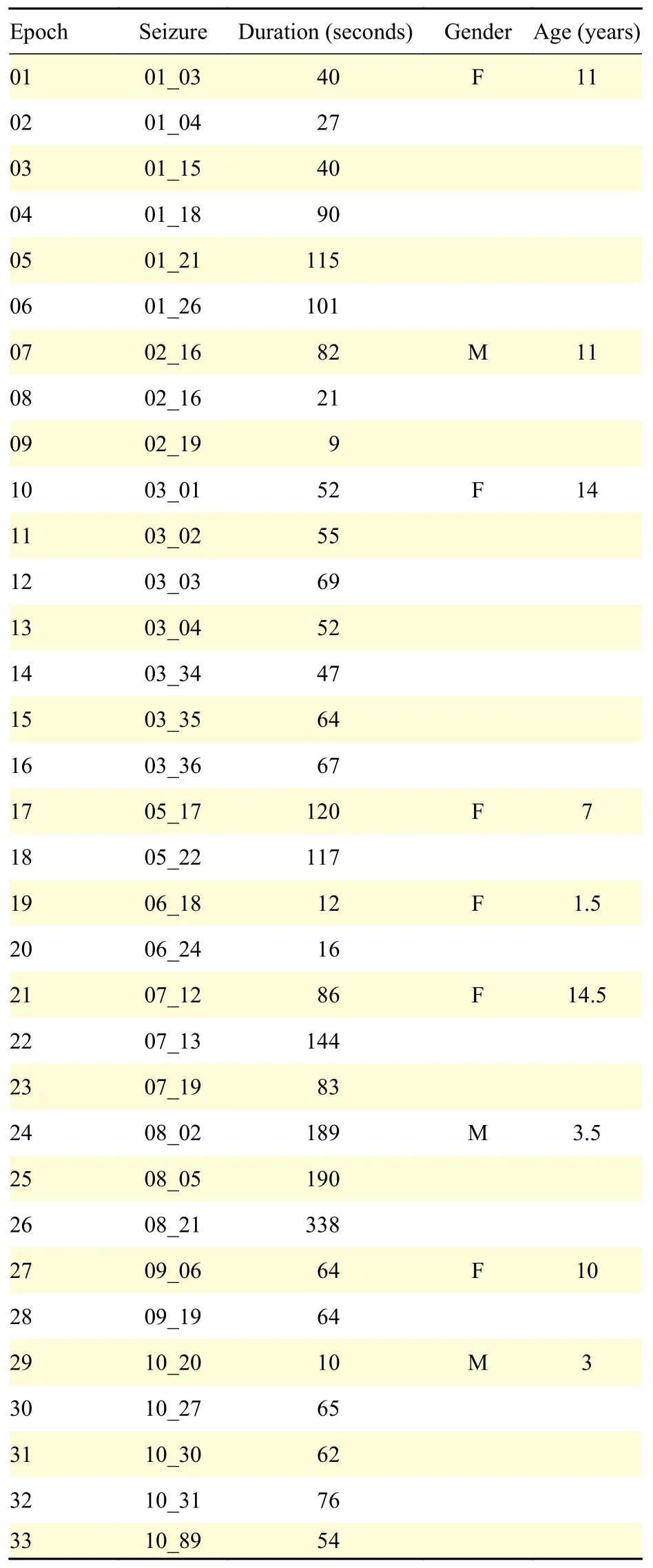
Table 1 Length of the 33 seizures used in this study
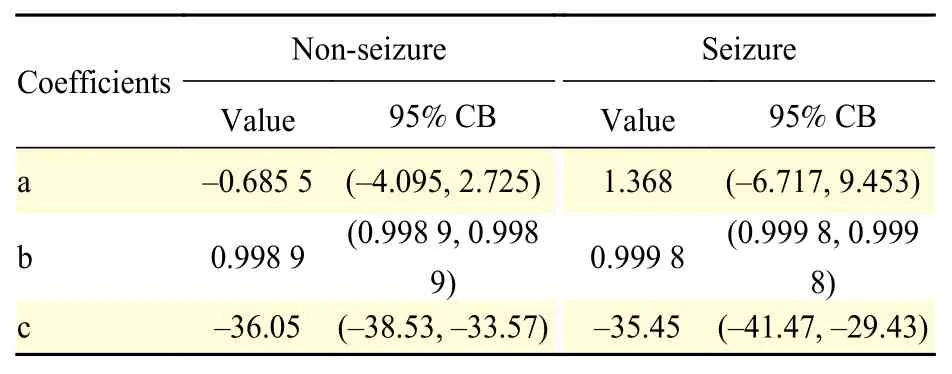
Table 2 The quadratic linear-parabolic model coefficients and the associated 95% confidence bounds (CB)
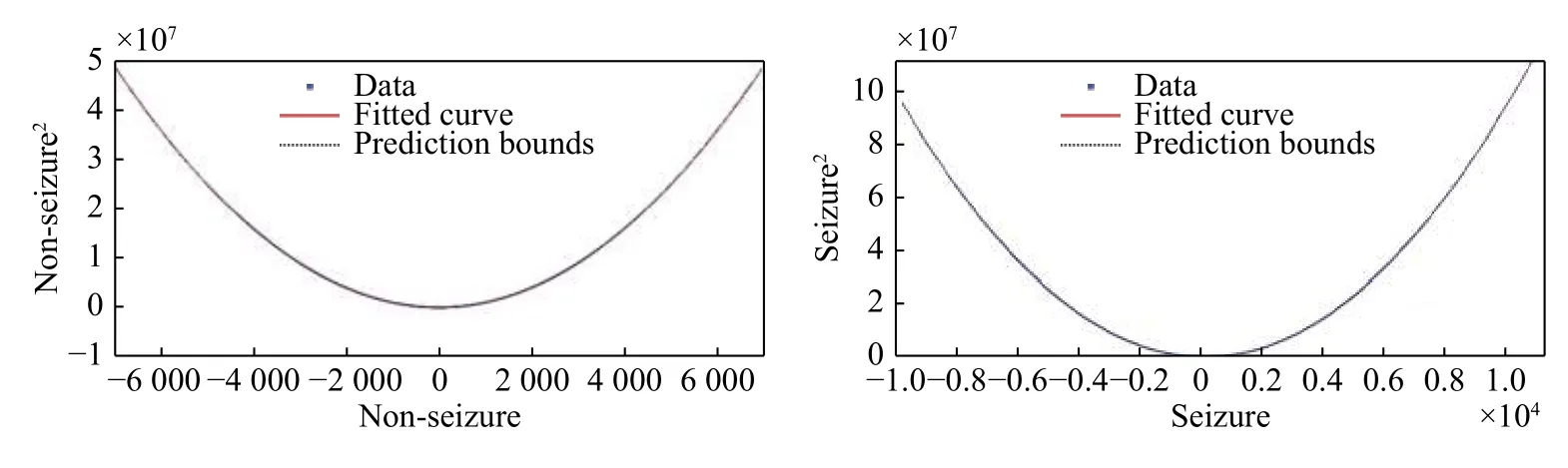
Fig. 1 Curves of the quadratic linear-parabolic model for seizure and non-seizure events. Note that the scale of the seizure curve(Right) is much larger than the one of the non-seizure curve (Left). The X-axis represents while the Y-axis represents (the unit on the X and Y axes correspond to the amplitude of the signal). The prediction bounds have a small uncertainty throughout the entire data range,therefore new observations can be predicted with high accuracy.

Fig. 2 EEG raw example. The onset detection begins in 81 seconds according to the medical annotation but by non-expert visual inspection cannot reach the same conclusion. The y-axis scale for each channel is between ±200 mv.
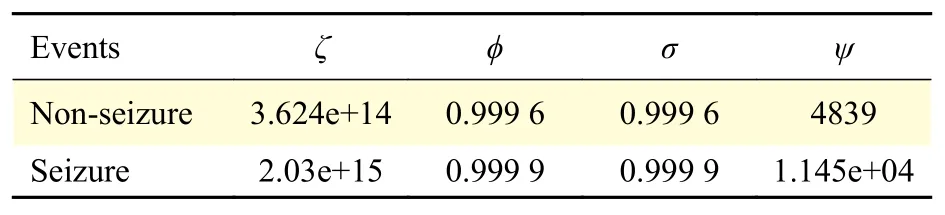
Table 3 Goodness-of-fit statistics
Estimating the four statistical parameters at each 1-second segment allows us to detect changes between epileptic EEG seizures. When we apply equation (7)to each 1-second segments, similar results are obtained as those applied to the signal as a whole.Fig. 3shows the similarity withFig. 1. Both seizure and non-seizure events have an excellent fitting with the linear-parabolic model, suggesting that the scale changes and the different values of the estimated parameters can be used to differentiate seizure and non-seizure events in EEG signals.
To cope with the high variability of EEG signals,the features were normalized to be in the range[0,1][33].Table 4reports the mean values for the four statistical parametersρ=[ζ,ϕ,σ,ψ]calculated for each 1-second segment fromX[n]. The weighted sum of squared residuals (ζ) has values closer to zero while R-square (ϕ) and adjusted R-square (σ) have values closer to one. This shows that the model has a high fit accuracy and can be used for seizure detection, as corroborated byFig. 3.
A value closer to zero in the weighted sum of squared residuals (ζ) and Root mean squared error (ψ)and values closer to one in R-square (ϕ) and adjusted R-squared (σ) suggest a good fit quality and therefore can be used as a predictor.
Fig. 4shows the main scatter plots of couples of the normalized statistical parameters (ζ,ϕ,σ,ψ). It is interesting to note that inζversusϕ(Fig. 4A) andψversusϕ(Fig. 4C) the seizure events (red dots) are concentrated closer to one with respect to non-seizure events (blue dots). InFig. 4B,ζversusψhas a big tendency to go to the right in the seizure events (red dots) with respect to non-seizure events (blue dots). InFig. 4D,ϕversusσshows a clear concentration of seizure events (red dots) with respect to non-seizure events (blue dots) on the upper right side. These scatter plots show that our predictor vectorρ=[ζ,ϕ,σ,ψ]is potentially useful to discriminate between seizures and non-seizures events in EEG signals.
To assess the performance of the proposed method,we adopted a supervised testing approach and used 1 518 events (1-second signals) to train and test the method with the 20-fold cross-validation technique of the predictor vectorρ=[ζ,ϕ,σ,ψ]. As explained above, the events were extracted from 33 seizure epochs and 33 non-seizure epochs, giving a total of 66 epochs from 9 different subjects. The method gives good classification performance, with 92% true positives rate (TPR) or sensitivity, 96% true negative rate (TNR) or specificity, 4% false positive rate, and 94.1% accuracy. Please note that segments for training and testing are drawn randomly without consideration of their epochs. This process was repeated 1 000 times to ensure not-bias in the partitioning.
Discussion
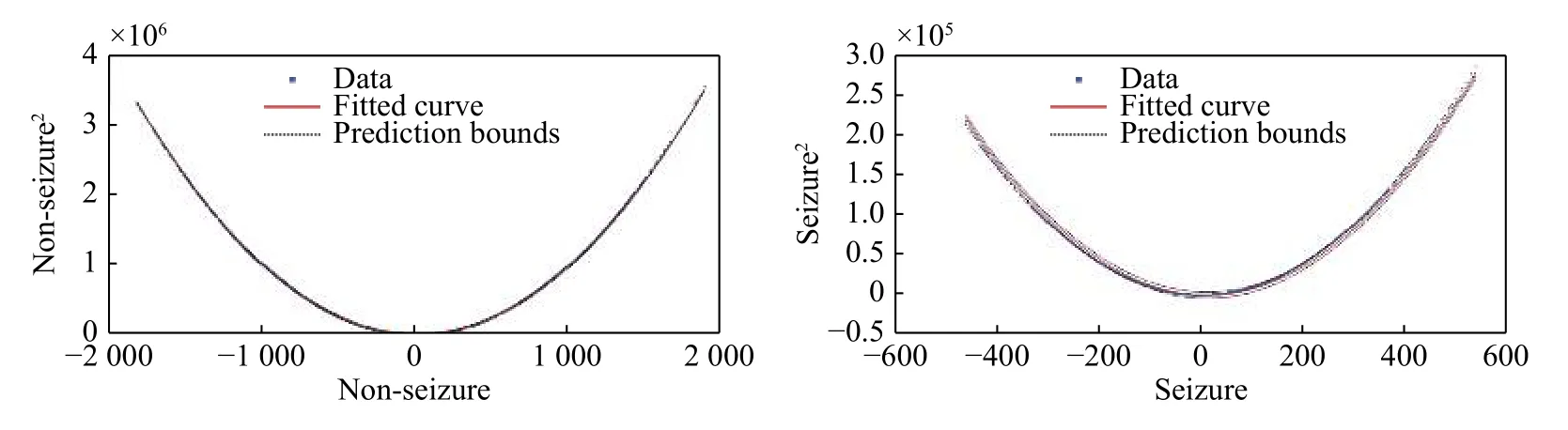
Fig. 3 Quadratic parabolic curve fitting example for one segment[n].
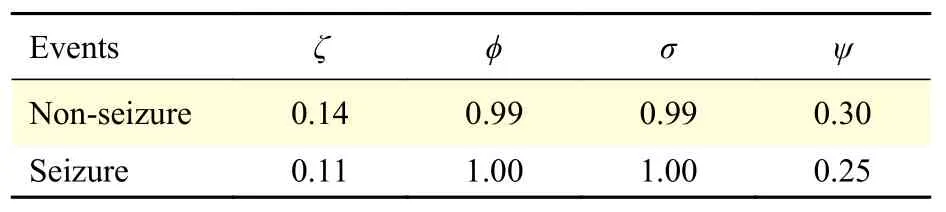
Table 4 Means of the statistical parameters with values normalized between 0 and 1 for all each 1-second time segments
This work presented a new method to detect epileptic seizures in EEG signals. The proposed model-based classification method relies on the design of a specific filter using the two-point central difference algorithm. A linear-parabolic model is fitted to the filtered signal using mean squares. Four statistical parameters associated with the model fitting are used as classification features to discriminate seizure and non-seizure events. A random forest classifier has been adopted to evaluate the ability of these parameters to detect seizures. The proposed methodology was applied to 66 balanced events from the Children Hospital Boston database. Results suggest that the proposed algorithm is a powerful tool for detecting epileptic seizure events in EEG signals achieving a 94.1% accuracy.
The proposed method has two main advantages compared to existing methods. First, its low computational complexity makes it implementable in quasi-real-time. Second, the method can be easily extended to work separately with different brain rhythms, by adapting the skip factorL. Despite these advantages, the method suffers some limitations,mainly noise and robustness.
The signal is affected by noise and artifacts due to the acquisition and pre-processing. This affects the precision of the results, especially when the method is generalized to complex epileptic forms. Outliers are not directly considered in the estimation procedure that needs more robustness.
Future work will focus on robust approaches to consider noise and artifacts. Optimization techniques will be investigated to remove outliers and improve the accuracy of the detection.
Acknowledgments
Part of the work was conducted by AQR during his Ph.D. work at Buenos Aires Institute of Technology(ITBA).
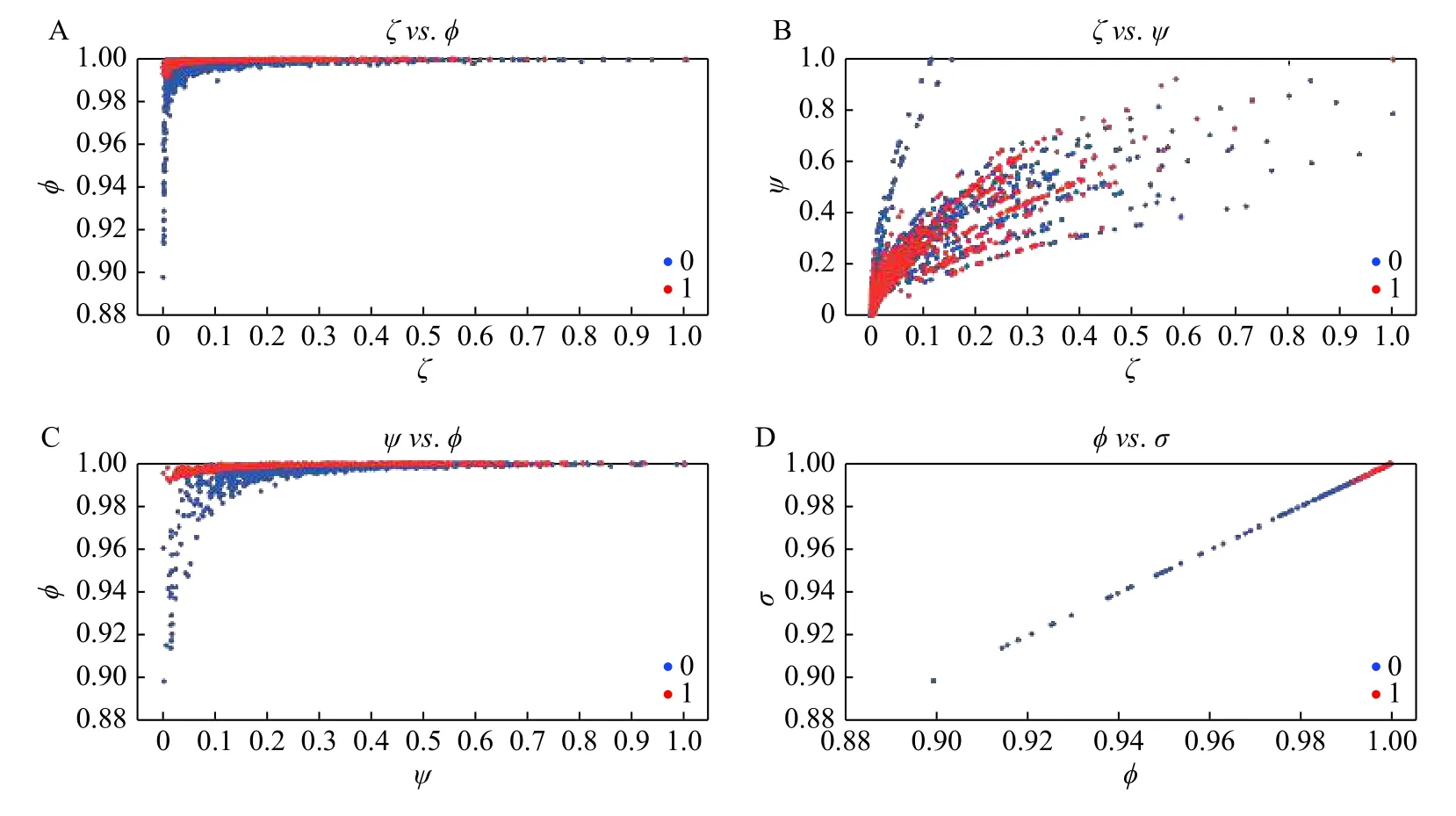
Fig. 4 Scatter plots for the four statistical parameters (ζ, ϕ, σ, ψ) observed through 1 518 events from 66 epochs, 33 seizures (red dots) and 33 on-seizures (blue dots) before the seizure. The weighted sum of squared residuals (ζ), R-square (ϕ), adjusted R-squared (σ)and root mean squared error (ψ) shows a clear concentration of seizure events (red dots) with respect to non-seizure events (blue dots).
杂志排行
THE JOURNAL OF BIOMEDICAL RESEARCH的其它文章
- Deep learning approach to detect seizure using reconstructed phase space images
- Epileptic seizure detection using EEG signals and extreme gradient boosting
- Complexity analysis and dynamic characteristics of EEG using MODWT based entropies for identification of seizure onset
- An optimized design of seizure detection system using joint feature extraction of multichannel EEG signals
- Classification of low-density EEG for epileptic seizures by energy and fractal features based on EMD
- Hidden Markov model based epileptic seizure detection using tunable Q wavelet transform
